Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CALU6 | SNV: p.G109R | . | |||
| FTC133 | SNV: p.V95M | . | |||
| KMS27 | SNV: p.V39L | DBIA Probe Info | |||
| MOLT4 | SNV: p.A2T | IA-alkyne Probe Info | |||
| SUDHL6 | SNV: p.R58Q | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
38.73 | LDD0324 | [1] | |
|
FBP2 Probe Info |
 |
37.85 | LDD0323 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K126(4.11) | LDD3494 | [3] | |
|
DBIA Probe Info |
 |
C53(5.51); C63(3.45) | LDD3311 | [4] | |
|
Alkyne-RA190 Probe Info |
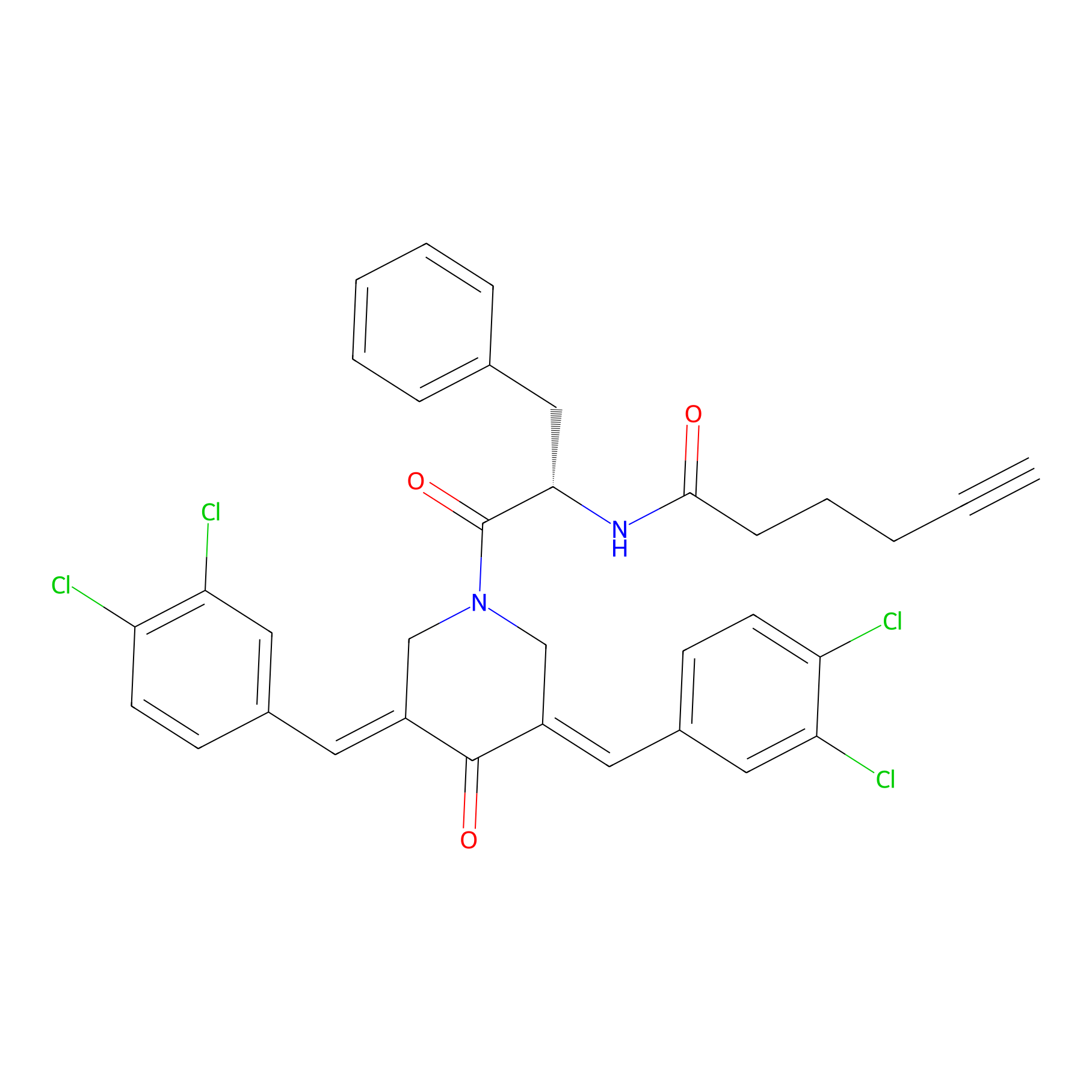 |
2.76 | LDD0299 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0166 | [7] | |
|
BTD Probe Info |
 |
N.A. | LDD0004 | [8] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
Acrolein Probe Info |
 |
H125(0.00); C9(0.00); C63(0.00) | LDD0217 | [10] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [10] | |
|
Methacrolein Probe Info |
 |
C63(0.00); C53(0.00) | LDD0218 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [11] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0139 | [11] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | H125(0.00); C63(0.00); C9(0.00) | LDD0222 | [10] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0022 | KB02 | A101D | C53(1.96) | LDD2250 | [4] |
| LDCM0023 | KB03 | A101D | C53(1.87) | LDD2667 | [4] |
| LDCM0024 | KB05 | G361 | C53(5.51); C63(3.45) | LDD3311 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [10] |
| LDCM0131 | RA190 | MM1.R | 2.76 | LDD0299 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Bone marrow stromal antigen 2 (BST2) | Tetherin family | Q10589 | |||
| CD81 antigen (CD81) | Tetraspanin (TM4SF) family | P60033 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Leukocyte immunoglobulin-like receptor subfamily A member 4 (LILRA4) | . | P59901 | |||
References
