Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
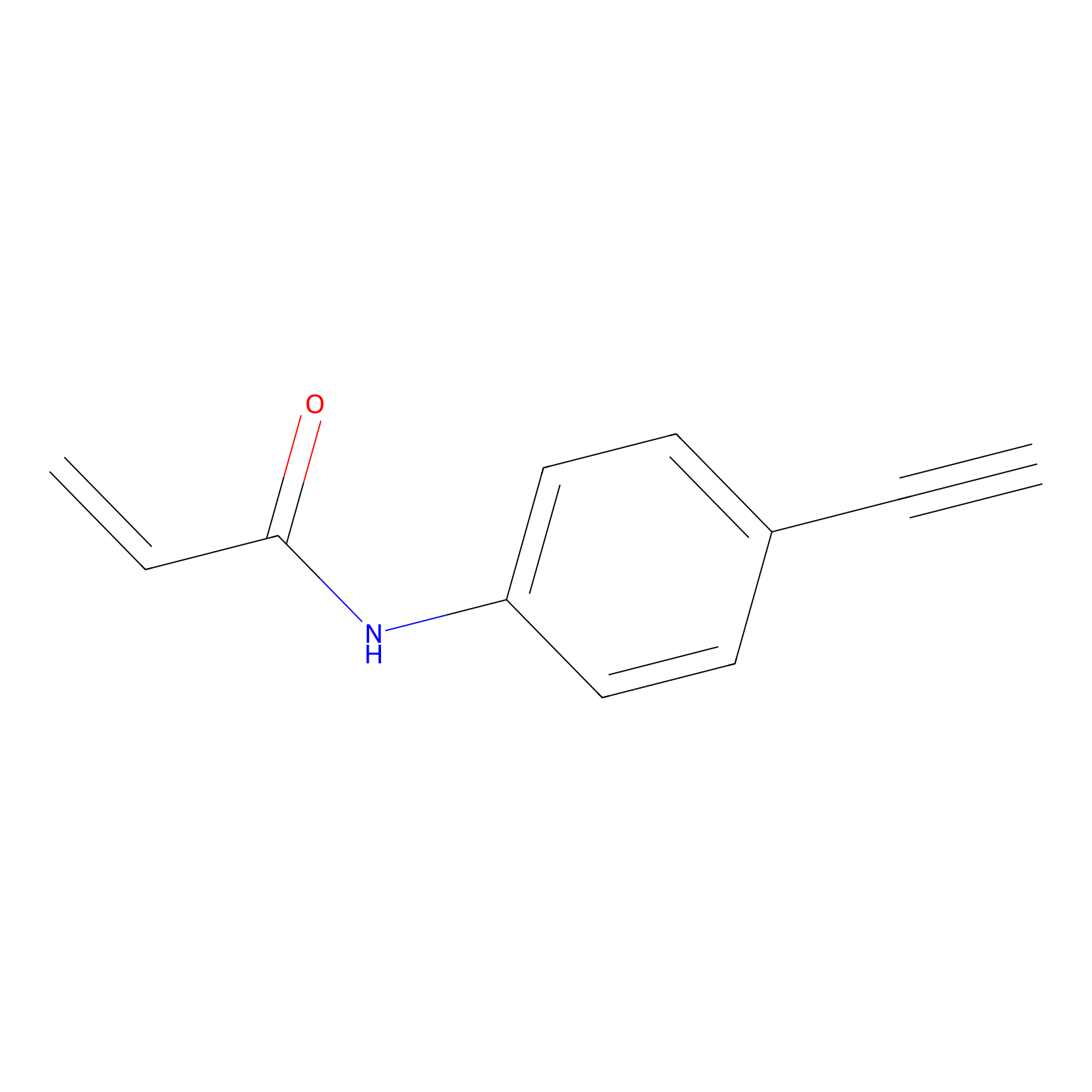 |
1.84 | LDD0453 | [1] | |
|
P8 Probe Info |
 |
1.54 | LDD0451 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
AZ-9 Probe Info |
 |
D184(0.99); E190(0.98) | LDD2208 | [3] | |
|
AF-1 Probe Info |
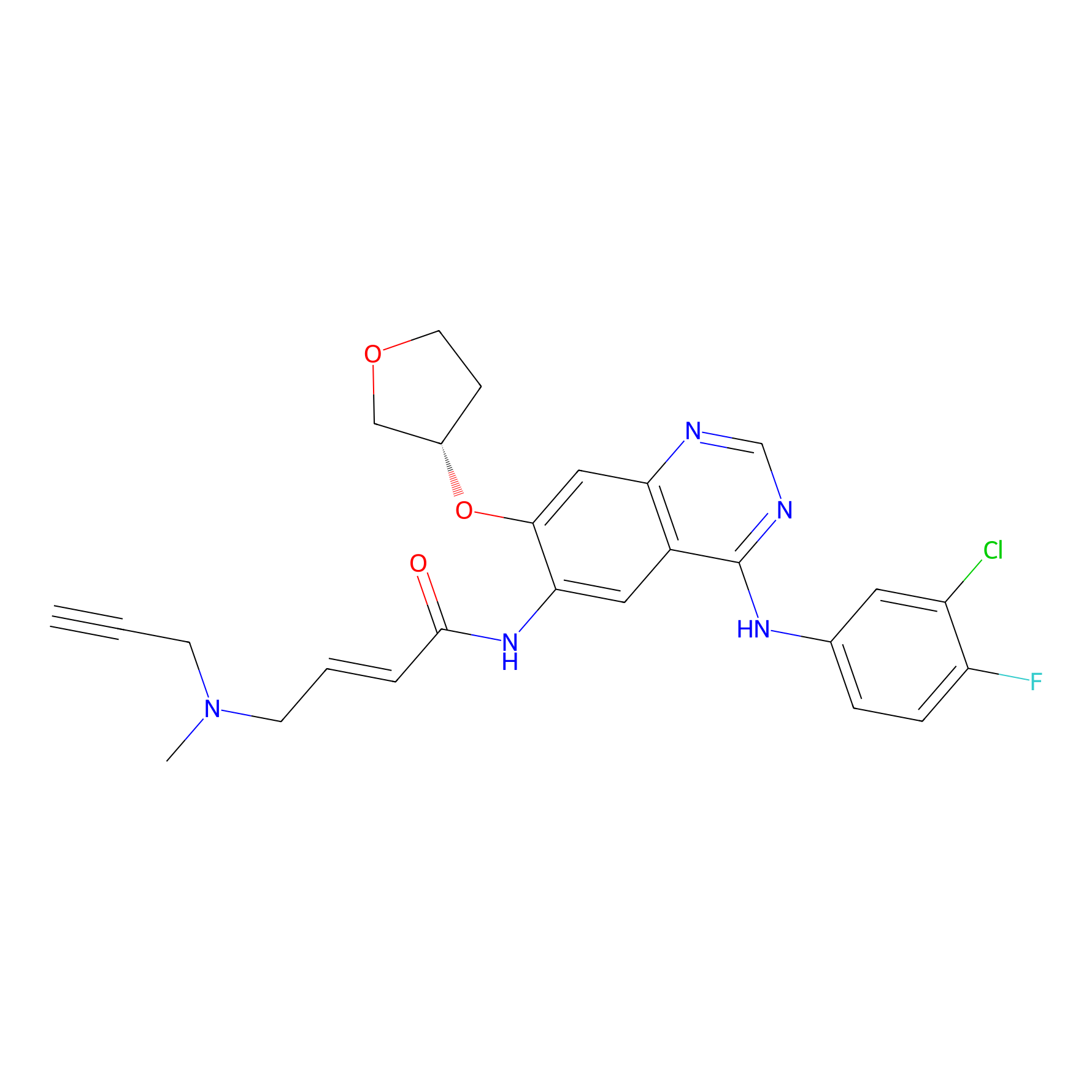 |
1.98 | LDD0421 | [4] | |
|
AF-2 Probe Info |
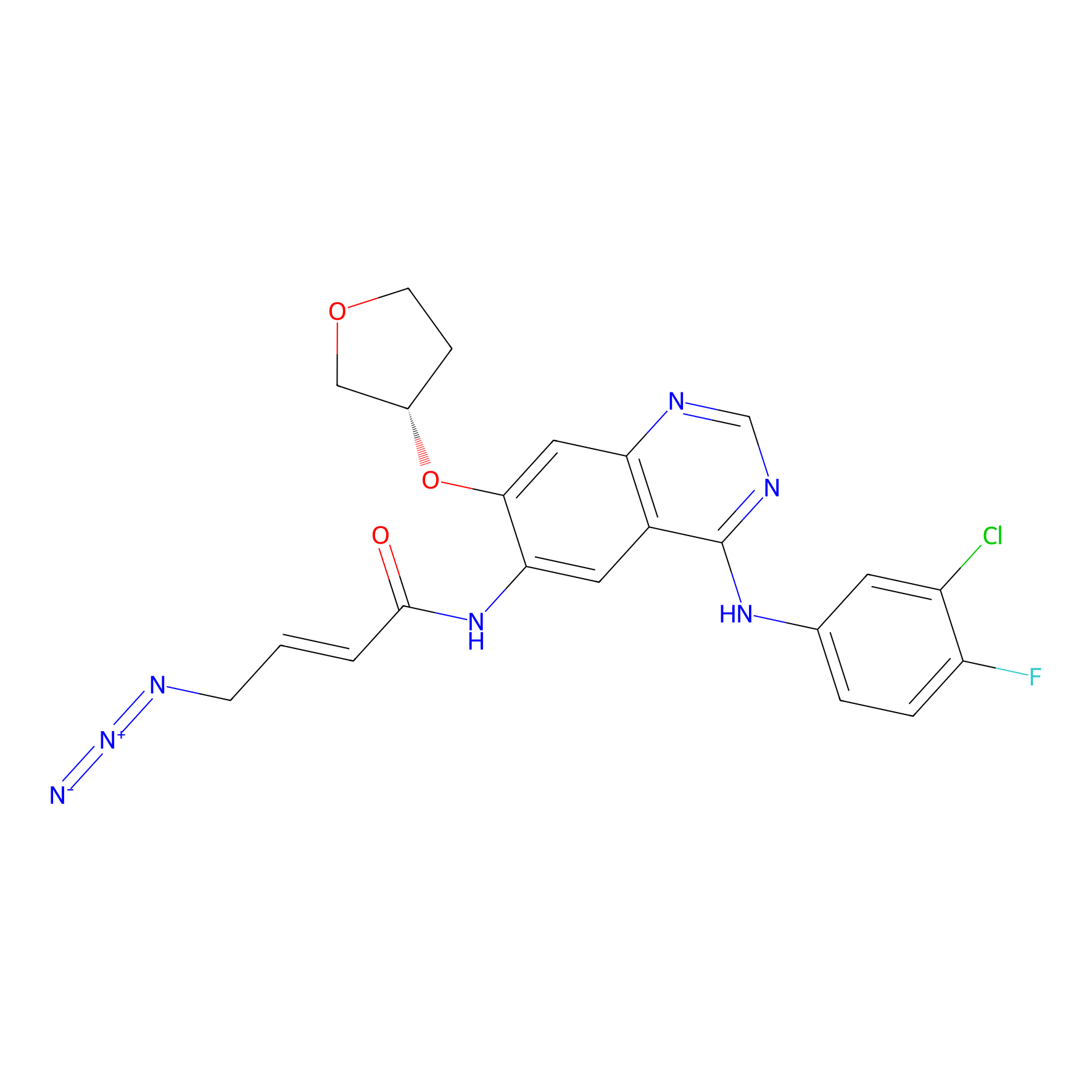 |
1.68 | LDD0422 | [4] | |
|
DBIA Probe Info |
 |
C494(5.31) | LDD3312 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0162 | [6] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
1.27 | LDD0136 | [8] | |
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [8] | |
|
VE-P Probe Info |
 |
N.A. | LDD0396 | [9] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Polyunsaturated fatty acid lipoxygenase ALOX12 (ALOX12) | Lipoxygenase family | P18054 | |||
| Kinesin-like protein KIFC3 (KIFC3) | Kinesin family | Q9BVG8 | |||
Other
References
