Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-5 Probe Info |
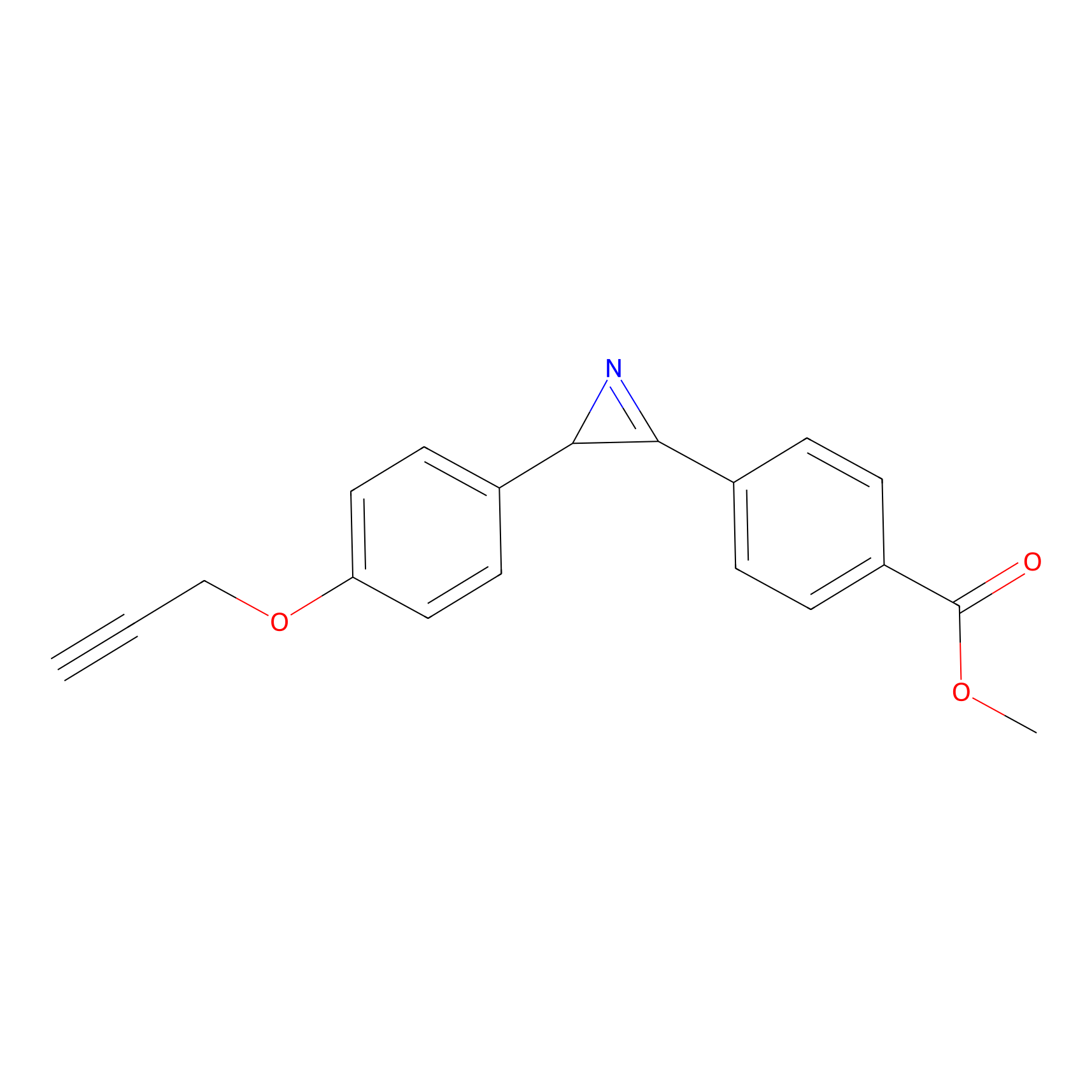 |
2.66 | LDD0394 | [1] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
ONAyne Probe Info |
 |
K296(0.68); K326(7.53); K353(4.99) | LDD0274 | [3] | |
|
STPyne Probe Info |
 |
K122(7.70); K130(3.16); K198(10.00); K199(8.01) | LDD0277 | [3] | |
|
AZ-9 Probe Info |
 |
E400(0.78); E128(1.30); D107(0.99); E153(10.00) | LDD2208 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K273(3.39); K296(3.61); K326(5.31); K331(5.69) | LDD3494 | [4] | |
|
P26 Probe Info |
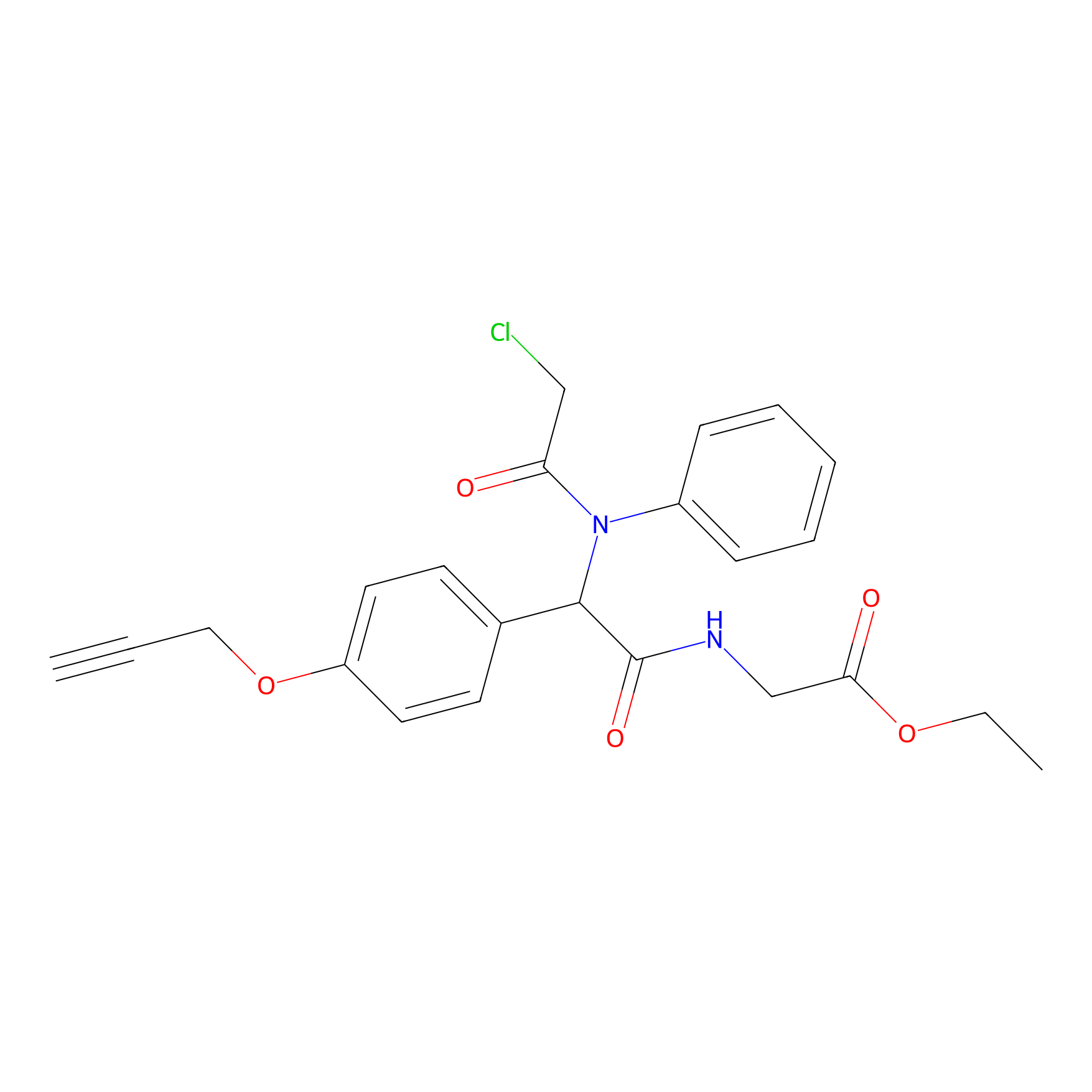 |
1.76 | LDD0409 | [5] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [6] | |
|
5E-2FA Probe Info |
 |
H297(0.00); H4(0.00) | LDD2235 | [7] | |
|
m-APA Probe Info |
 |
H297(0.00); H4(0.00) | LDD2231 | [7] | |
|
ATP probe Probe Info |
 |
K296(0.00); K331(0.00); K108(0.00); K96(0.00) | LDD0035 | [8] | |
|
HHS-465 Probe Info |
 |
K394(0.00); Y55(0.00); K117(0.00); K108(0.00) | LDD2240 | [9] | |
|
HHS-475 Probe Info |
 |
Y268(0.71); Y55(0.75) | LDD2238 | [10] | |
|
HHS-482 Probe Info |
 |
Y205(0.56); Y268(0.60); Y283(0.61); Y40(0.33) | LDD2239 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
3.41 | LDD0136 | [11] | |
|
STS-2 Probe Info |
 |
2.02 | LDD0138 | [11] | |
|
DA-2 Probe Info |
 |
N.A. | LDD0071 | [12] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
