Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
W1 Probe Info |
 |
75.43 | LDD0235 | [1] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [2] | |
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [2] | |
|
IPM Probe Info |
 |
C469(0.00); C350(0.00); C656(0.00); C503(0.00) | LDD0241 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K467(0.07); K135(1.26); K134(2.36) | LDD3494 | [3] | |
|
DBIA Probe Info |
 |
C469(1.01) | LDD3314 | [4] | |
|
BTD Probe Info |
 |
C469(1.29) | LDD2101 | [5] | |
|
YY4-yne Probe Info |
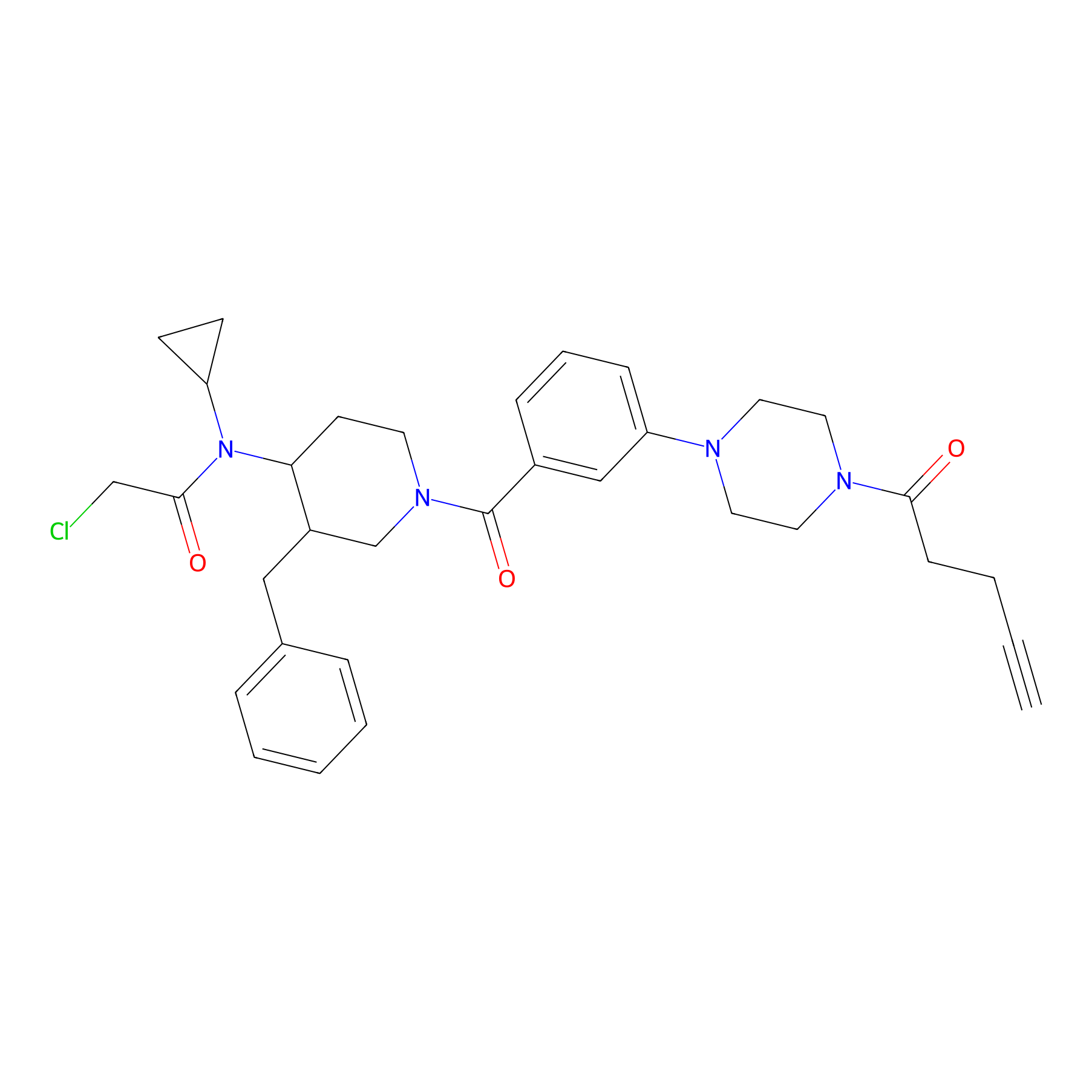 |
3.57 | LDD0400 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [8] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [10] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alk-rapa Probe Info |
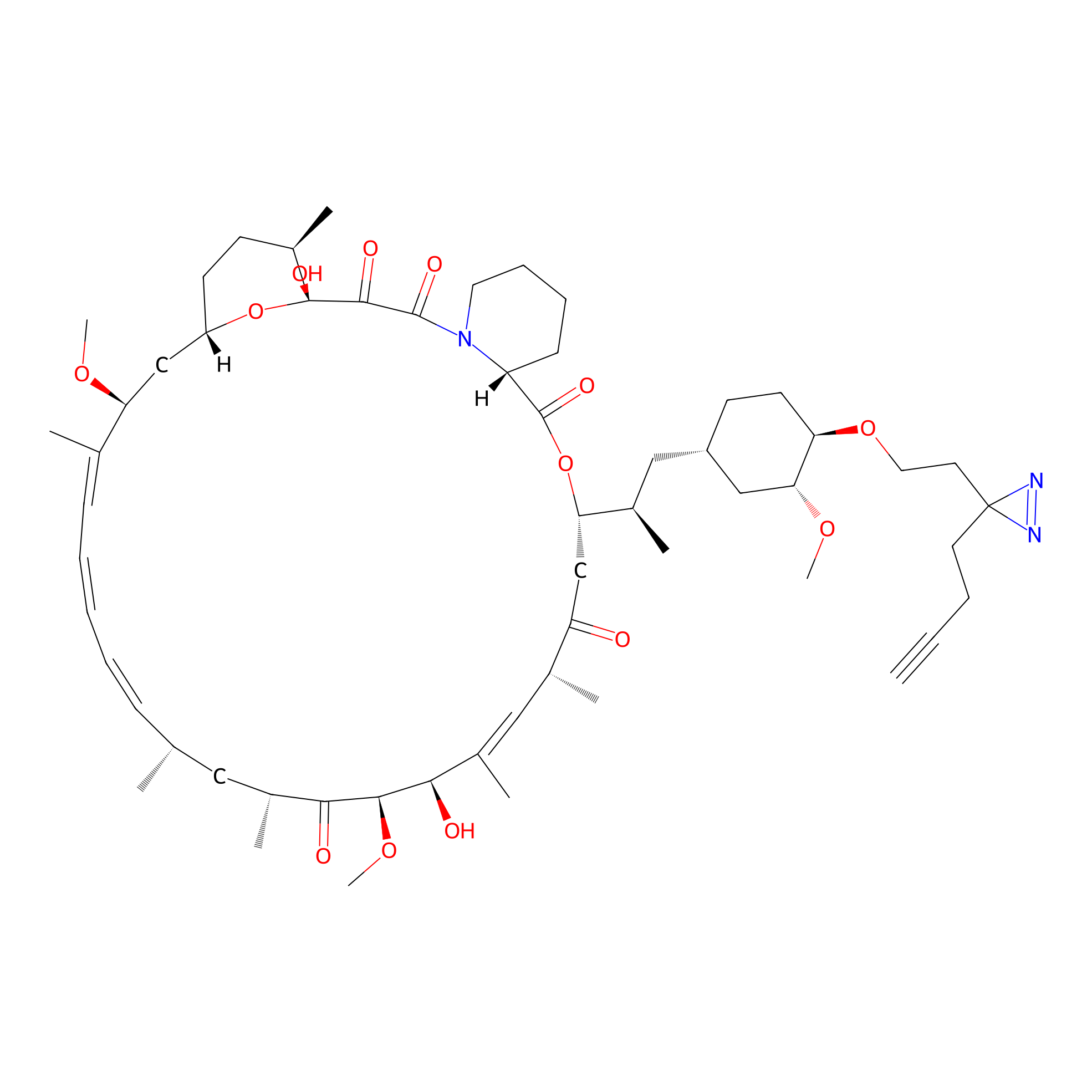 |
2.67 | LDD0213 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0215 | AC10 | HEK-293T | C137(0.85) | LDD1508 | [12] |
| LDCM0277 | AC18 | HEK-293T | C137(0.89) | LDD1516 | [12] |
| LDCM0279 | AC2 | HEK-293T | C137(1.02) | LDD1518 | [12] |
| LDCM0286 | AC26 | HEK-293T | C137(0.83) | LDD1525 | [12] |
| LDCM0295 | AC34 | HEK-293T | C137(1.46) | LDD1534 | [12] |
| LDCM0304 | AC42 | HEK-293T | C137(1.16) | LDD1543 | [12] |
| LDCM0313 | AC50 | HEK-293T | C137(0.86) | LDD1552 | [12] |
| LDCM0321 | AC58 | HEK-293T | C137(1.00) | LDD1560 | [12] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0367 | CL1 | HEK-293T | C137(0.85) | LDD1571 | [12] |
| LDCM0370 | CL101 | HEK-293T | C137(0.95) | LDD1574 | [12] |
| LDCM0374 | CL105 | HEK-293T | C137(1.03) | LDD1578 | [12] |
| LDCM0378 | CL109 | HEK-293T | C137(1.02) | LDD1582 | [12] |
| LDCM0383 | CL113 | HEK-293T | C137(1.09) | LDD1587 | [12] |
| LDCM0387 | CL117 | HEK-293T | C137(1.11) | LDD1591 | [12] |
| LDCM0392 | CL121 | HEK-293T | C137(1.01) | LDD1596 | [12] |
| LDCM0396 | CL125 | HEK-293T | C137(0.82) | LDD1600 | [12] |
| LDCM0400 | CL13 | HEK-293T | C137(1.00) | LDD1604 | [12] |
| LDCM0405 | CL18 | HEK-293T | C137(0.96) | LDD1609 | [12] |
| LDCM0413 | CL25 | HEK-293T | C137(0.97) | LDD1617 | [12] |
| LDCM0419 | CL30 | HEK-293T | C137(0.93) | LDD1623 | [12] |
| LDCM0426 | CL37 | HEK-293T | C137(0.93) | LDD1630 | [12] |
| LDCM0432 | CL42 | HEK-293T | C137(0.84) | LDD1636 | [12] |
| LDCM0439 | CL49 | HEK-293T | C137(0.96) | LDD1643 | [12] |
| LDCM0445 | CL54 | HEK-293T | C137(0.99) | LDD1648 | [12] |
| LDCM0451 | CL6 | HEK-293T | C137(0.91) | LDD1654 | [12] |
| LDCM0453 | CL61 | HEK-293T | C137(0.97) | LDD1656 | [12] |
| LDCM0458 | CL66 | HEK-293T | C137(1.03) | LDD1661 | [12] |
| LDCM0466 | CL73 | HEK-293T | C137(1.03) | LDD1669 | [12] |
| LDCM0471 | CL78 | HEK-293T | C137(1.01) | LDD1674 | [12] |
| LDCM0479 | CL85 | HEK-293T | C137(0.78) | LDD1682 | [12] |
| LDCM0485 | CL90 | HEK-293T | C137(1.27) | LDD1688 | [12] |
| LDCM0492 | CL97 | HEK-293T | C137(1.03) | LDD1695 | [12] |
| LDCM0022 | KB02 | 769-P | C469(1.14) | LDD2246 | [4] |
| LDCM0023 | KB03 | A101D | C596(4.28) | LDD2667 | [4] |
| LDCM0024 | KB05 | IGR37 | C469(1.01) | LDD3314 | [4] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C469(1.29) | LDD2101 | [5] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C469(0.96) | LDD2111 | [5] |
| LDCM0090 | Rapamycin | JHH-7 | 2.67 | LDD0213 | [11] |
| LDCM0113 | W17 | Hep-G2 | C469(6.47) | LDD0240 | [1] |
| LDCM0154 | YY4 | T cell | 3.57 | LDD0400 | [6] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Paired-like homeodomain transcription factor LEUTX (LEUTX) | Paired homeobox family | A8MZ59 | |||
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| G-protein coupled receptor 42 (GPR42) | G-protein coupled receptor 1 family | O15529 | |||
Other
The Drug(s) Related To This Target
Approved
Phase 1
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Calaa-01 | . | D0A1GR | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Ep-2167 | . | D0QB8Q | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Chromic Citrate | . | DB14526 | |||
| Dichlorvos | . | DB11397 | |||
References
