Details of the Target
General Information of Target
| Target ID | LDTP14895 | |||||
|---|---|---|---|---|---|---|
| Target Name | Eukaryotic translation initiation factor 2 subunit 3B (EIF2S3B) | |||||
| Gene Name | EIF2S3B | |||||
| Synonyms |
Eukaryotic translation initiation factor 2 subunit 3B; EC 3.6.5.3; Eukaryotic translation initiation factor 2 subunit gamma A; eIF-2-gamma A; eIF-2gA |
|||||
| 3D Structure | ||||||
| Sequence |
MSPQGPAVLSLGSLCLDTNQAPNWTGLQTLLQQLPPQDIDERYCLALGEEERAELQLFCA
RRKQEALGQGVARLVLPKLEGHTCEKCRELLKPGEYGVFAARAGEQRCWHQPCFACQACG QALINLIYFYHDGQLYCGRHHAELLRPRCPACDQLIFSWRCTEAEGQRWHENHFCCQDCA GPLGGGRYALPGGSPCCPSCFENRYSDAGSSWAGALEGQAFLGETGLDRTEGRDQTSVNS ATLSRTLLAAAGGSSLQTQRGLPGSSPQQENRPGDKAEAPKGQEQCRLETIRDPKDTPFS TCSSSSDSEPEGFFLGERLPQSWKTPGSLQAEDSNASKTHCTMC |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
TRAFAC class translation factor GTPase superfamily, Classic translation factor GTPase family, EIF2G subfamily
|
|||||
| Function |
Member of the eIF2 complex that functions in the early steps of protein synthesis by forming a ternary complex with GTP and initiator tRNA. This complex binds to a 40S ribosomal subunit, followed by mRNA binding to form the 43S pre-initiation complex (43S PIC). Junction of the 60S ribosomal subunit to form the 80S initiation complex is preceded by hydrolysis of the GTP bound to eIF2 and release of an eIF2-GDP binary complex. In order for eIF2 to recycle and catalyze another round of initiation, the GDP bound to eIF2 must exchange with GTP by way of a reaction catalyzed by eIF-2B.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | SNV: p.L158M | DBIA Probe Info | |||
| COLO678 | Insertion: p.E5GfsTer11 | DBIA Probe Info | |||
| COLO679 | SNV: p.G409R | DBIA Probe Info | |||
| CORL88 | SNV: p.Q329Ter | DBIA Probe Info | |||
| HEC1 | SNV: p.S225L | DBIA Probe Info | |||
| HUH1 | SNV: p.L371S | DBIA Probe Info | |||
| NCIH2291 | Insertion: p.V319CfsTer8 | DBIA Probe Info | |||
| PEO1 | Deletion: p.P345_D351del | DBIA Probe Info | |||
| SKOV3 | SNV: p.R460S | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y330(5.65) | LDD0257 | [1] | |
|
AZ-9 Probe Info |
 |
E5(10.00) | LDD2209 | [2] | |
|
DBIA Probe Info |
 |
C348(2.82); C236(1.30) | LDD3310 | [3] | |
|
BTD Probe Info |
 |
C348(1.66) | LDD1700 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C269(2.03) | LDD0169 | [5] | |
|
IPM Probe Info |
 |
C96(0.52) | LDD1701 | [4] | |
|
Acrolein Probe Info |
 |
C269(0.00); C434(0.00); H325(0.00); H184(0.00) | LDD0217 | [6] | |
|
Cinnamaldehyde Probe Info |
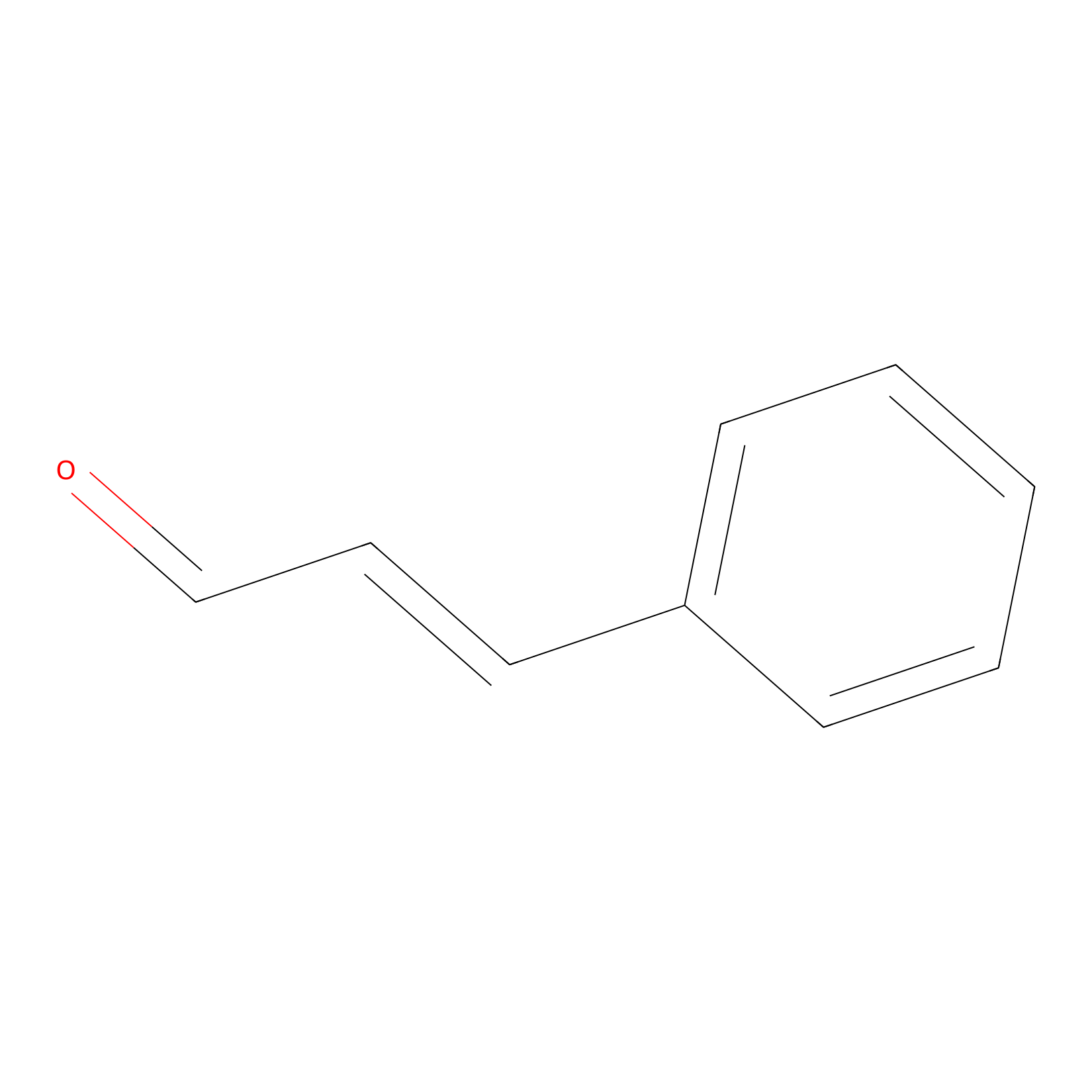 |
C348(0.00); C434(0.00) | LDD0220 | [6] | |
|
Crotonaldehyde Probe Info |
 |
C434(0.00); C348(0.00) | LDD0219 | [6] | |
|
Methacrolein Probe Info |
 |
C434(0.00); C269(0.00); C348(0.00); C101(0.00) | LDD0218 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-1 Probe Info |
 |
N.A. | LDD0137 | [7] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C348(0.39) | LDD2142 | [4] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C348(0.67); C269(0.65) | LDD2112 | [4] |
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C434(0.87); C269(0.95); C101(1.09); C96(0.64) | LDD2117 | [4] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C434(1.41); C348(1.00) | LDD2152 | [4] |
| LDCM0538 | 4-(Cyanoacetyl)morpholine | MDA-MB-231 | C348(1.58) | LDD2131 | [4] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C269(2.03) | LDD0169 | [5] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C348(0.69); C96(0.69) | LDD2113 | [4] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C434(0.62); C269(0.84) | LDD2091 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C269(0.00); C434(0.00); C348(0.00); H33(0.00) | LDD0222 | [6] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C96(0.66) | LDD1702 | [4] |
| LDCM0107 | IAA | HeLa | C269(0.00); H33(0.00); C434(0.00); H184(0.00) | LDD0221 | [6] |
| LDCM0022 | KB02 | 22RV1 | C236(1.09) | LDD2243 | [3] |
| LDCM0023 | KB03 | MDA-MB-231 | C96(0.52) | LDD1701 | [4] |
| LDCM0024 | KB05 | COLO792 | C348(2.82); C236(1.30) | LDD3310 | [3] |
| LDCM0109 | NEM | HeLa | H325(0.00); H33(0.00); H184(0.00); H65(0.00) | LDD0223 | [6] |
| LDCM0496 | Nucleophilic fragment 11a | MDA-MB-231 | C348(0.96) | LDD2089 | [4] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C348(1.18) | LDD2090 | [4] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C434(0.88); C348(0.92) | LDD2093 | [4] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C434(1.01); C348(0.64); C269(1.00) | LDD2097 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C348(0.95); C269(0.95) | LDD2099 | [4] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C348(0.47); C269(0.41) | LDD2100 | [4] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C434(0.86); C348(0.53); C269(0.77); C101(1.53) | LDD2101 | [4] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C348(0.45); C269(0.31) | LDD2104 | [4] |
| LDCM0513 | Nucleophilic fragment 19b | MDA-MB-231 | C269(0.15) | LDD2106 | [4] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C434(0.94); C348(0.97); C269(1.01) | LDD2107 | [4] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C434(0.51); C348(0.48) | LDD2108 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C434(0.47); C348(0.52); C269(0.75); C96(0.50) | LDD2109 | [4] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C348(0.51) | LDD2110 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C434(1.05) | LDD2111 | [4] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C348(0.69); C269(0.36) | LDD2114 | [4] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C348(0.38) | LDD2115 | [4] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C434(1.87); C348(1.76) | LDD2119 | [4] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C269(0.82) | LDD2120 | [4] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C434(0.75); C348(0.83); C269(1.03); C101(0.82) | LDD2123 | [4] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C348(0.74); C269(0.86); C101(0.74) | LDD2125 | [4] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C434(1.02); C348(0.89); C269(1.51); C101(0.82) | LDD2127 | [4] |
| LDCM0535 | Nucleophilic fragment 30b | MDA-MB-231 | C269(0.52) | LDD2128 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C434(1.23); C348(1.02) | LDD2129 | [4] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C348(0.39) | LDD2133 | [4] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C348(0.30); C101(0.81) | LDD2134 | [4] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C434(1.54); C348(1.30) | LDD2135 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C434(1.06); C269(1.18); C101(1.16); C96(1.09) | LDD2136 | [4] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C434(1.01); C348(0.92); C269(0.94); C96(0.81) | LDD2137 | [4] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C348(1.66) | LDD1700 | [4] |
| LDCM0546 | Nucleophilic fragment 40 | MDA-MB-231 | C348(0.75); C269(1.06) | LDD2140 | [4] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C269(0.53) | LDD2141 | [4] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C269(0.62) | LDD2143 | [4] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C434(2.53); C348(1.95); C269(2.42) | LDD2144 | [4] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C348(0.41) | LDD2145 | [4] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C434(0.93); C348(0.85) | LDD2146 | [4] |
| LDCM0553 | Nucleophilic fragment 6b | MDA-MB-231 | C269(1.57) | LDD2147 | [4] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C434(0.38); C348(0.34); C269(0.46); C101(0.59) | LDD2148 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C434(0.44); C269(0.56) | LDD2150 | [4] |
References
