Details of the Target
General Information of Target
| Target ID | LDTP13034 | |||||
|---|---|---|---|---|---|---|
| Target Name | Prostaglandin F2 receptor negative regulator (PTGFRN) | |||||
| Gene Name | PTGFRN | |||||
| Gene ID | 5738 | |||||
| Synonyms |
CD9P1; EWIF; FPRP; KIAA1436; Prostaglandin F2 receptor negative regulator; CD9 partner 1; CD9P-1; Glu-Trp-Ile EWI motif-containing protein F; EWI-F; Prostaglandin F2-alpha receptor regulatory protein; Prostaglandin F2-alpha receptor-associated protein; CD antigen CD315
|
|||||
| 3D Structure | ||||||
| Sequence |
MAELQQLQEFEIPTGREALRGNHSALLRVADYCEDNYVQATDKRKALEETMAFTTQALAS
VAYQVGNLAGHTLRMLDLQGAALRQVEARVSTLGQMVNMHMEKVARREIGTLATVQRLPP GQKVIAPENLPPLTPYCRRPLNFGCLDDIGHGIKDLSTQLSRTGTLSRKSIKAPATPASA TLGRPPRIPEPVHLPVVPDGRLSAASSAFSLASAGSAEGVGGAPTPKGQAAPPAPPLPSS LDPPPPPAAVEVFQRPPTLEELSPPPPDEELPLPLDLPPPPPLDGDELGLPPPPPGFGPD EPSWVPASYLEKVVTLYPYTSQKDNELSFSEGTVICVTRRYSDGWCEGVSSEGTGFFPGN YVEPSC |
|||||
| Target Bioclass |
Immunoglobulin
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function |
Inhibits the binding of prostaglandin F2-alpha (PGF2-alpha) to its specific FP receptor, by decreasing the receptor number rather than the affinity constant. Functional coupling with the prostaglandin F2-alpha receptor seems to occur. In myoblasts, associates with tetraspanins CD9 and CD81 to prevent myotube fusion during muscle regeneration.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
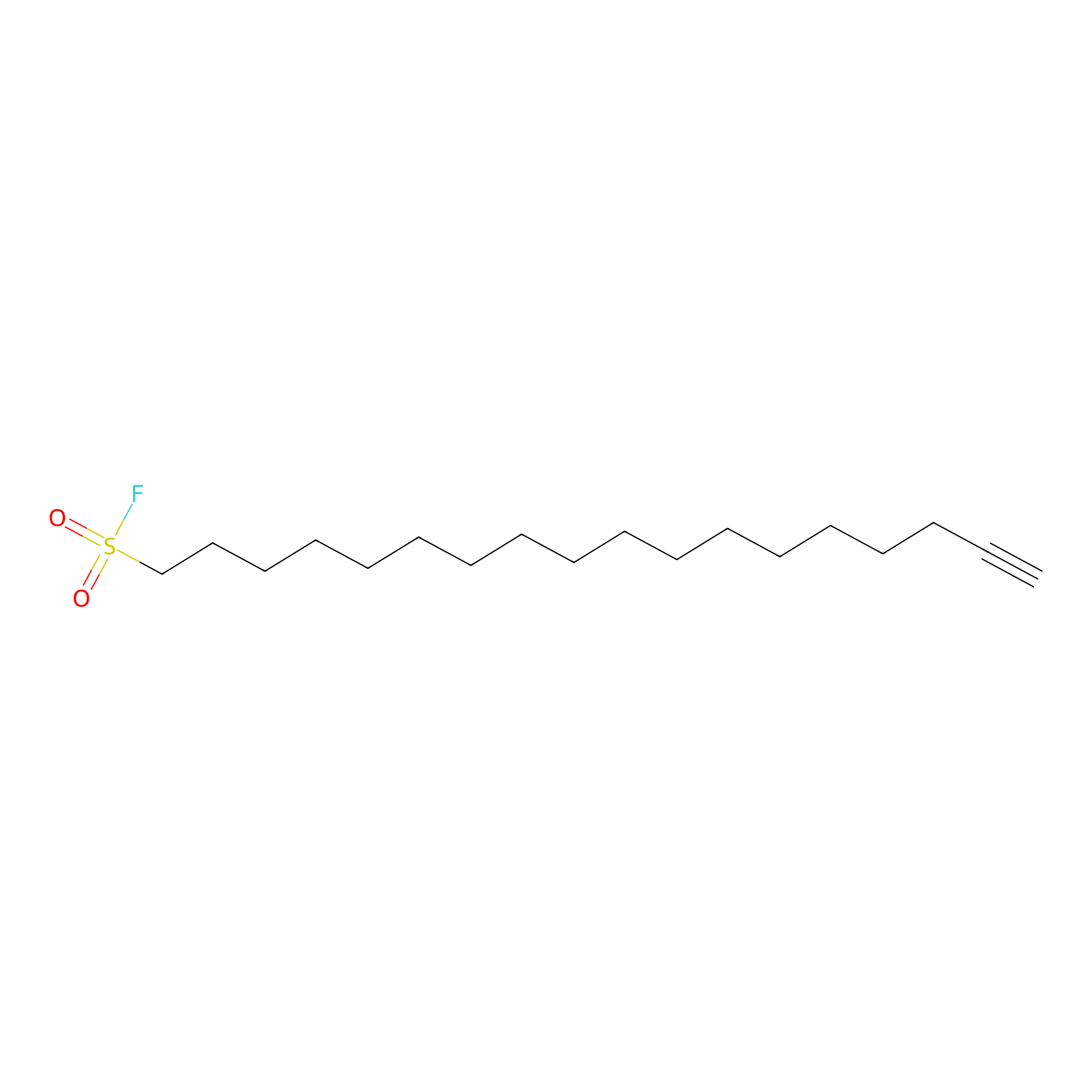 |
1.85 | LDD0197 | [1] | |
|
FBPP2 Probe Info |
 |
19.41 | LDD0318 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K492(0.47); K522(2.13); K579(2.67); K576(2.67) | LDD3494 | [3] | |
|
DBIA Probe Info |
 |
C429(2.42) | LDD3314 | [4] | |
|
5E-2FA Probe Info |
 |
H206(0.00); H146(0.00) | LDD2235 | [5] | |
|
m-APA Probe Info |
 |
H206(0.00); H146(0.00) | LDD2231 | [5] | |
|
IA-alkyne Probe Info |
 |
C169(0.00); C655(0.00); C429(0.00) | LDD0165 | [6] | |
|
Acrolein Probe Info |
 |
H350(0.00); H116(0.00) | LDD0217 | [7] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C299(0.92) | LDD1507 | [8] |
| LDCM0237 | AC12 | HEK-293T | C299(1.00) | LDD1510 | [8] |
| LDCM0259 | AC14 | HEK-293T | C299(1.00); C119(0.58) | LDD1512 | [8] |
| LDCM0276 | AC17 | HEK-293T | C299(1.04) | LDD1515 | [8] |
| LDCM0280 | AC20 | HEK-293T | C299(0.97) | LDD1519 | [8] |
| LDCM0281 | AC21 | HEK-293T | C299(0.99) | LDD1520 | [8] |
| LDCM0282 | AC22 | HEK-293T | C299(0.80); C119(0.73) | LDD1521 | [8] |
| LDCM0285 | AC25 | HEK-293T | C299(1.05) | LDD1524 | [8] |
| LDCM0288 | AC28 | HEK-293T | C299(0.88) | LDD1527 | [8] |
| LDCM0289 | AC29 | HEK-293T | C299(1.07) | LDD1528 | [8] |
| LDCM0291 | AC30 | HEK-293T | C299(0.73); C119(0.90) | LDD1530 | [8] |
| LDCM0294 | AC33 | HEK-293T | C299(1.23) | LDD1533 | [8] |
| LDCM0297 | AC36 | HEK-293T | C299(1.02) | LDD1536 | [8] |
| LDCM0298 | AC37 | HEK-293T | C299(0.90) | LDD1537 | [8] |
| LDCM0299 | AC38 | HEK-293T | C299(0.82); C119(0.68) | LDD1538 | [8] |
| LDCM0301 | AC4 | HEK-293T | C299(0.92) | LDD1540 | [8] |
| LDCM0303 | AC41 | HEK-293T | C299(0.80) | LDD1542 | [8] |
| LDCM0306 | AC44 | HEK-293T | C299(1.01) | LDD1545 | [8] |
| LDCM0307 | AC45 | HEK-293T | C299(1.07) | LDD1546 | [8] |
| LDCM0308 | AC46 | HEK-293T | C299(0.94); C119(0.91) | LDD1547 | [8] |
| LDCM0311 | AC49 | HEK-293T | C299(0.94) | LDD1550 | [8] |
| LDCM0312 | AC5 | HEK-293T | C299(0.93) | LDD1551 | [8] |
| LDCM0315 | AC52 | HEK-293T | C299(0.92) | LDD1554 | [8] |
| LDCM0316 | AC53 | HEK-293T | C299(1.02) | LDD1555 | [8] |
| LDCM0317 | AC54 | HEK-293T | C299(0.97); C119(0.65) | LDD1556 | [8] |
| LDCM0320 | AC57 | HEK-293T | C299(1.08) | LDD1559 | [8] |
| LDCM0323 | AC6 | HEK-293T | C299(0.90); C119(0.87) | LDD1562 | [8] |
| LDCM0324 | AC60 | HEK-293T | C299(0.91) | LDD1563 | [8] |
| LDCM0325 | AC61 | HEK-293T | C299(0.89) | LDD1564 | [8] |
| LDCM0326 | AC62 | HEK-293T | C299(0.85); C119(0.65) | LDD1565 | [8] |
| LDCM0248 | AKOS034007472 | HEK-293T | C299(1.05) | LDD1511 | [8] |
| LDCM0356 | AKOS034007680 | HEK-293T | C299(0.93) | LDD1570 | [8] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [7] |
| LDCM0368 | CL10 | HEK-293T | C299(0.96); C119(0.70) | LDD1572 | [8] |
| LDCM0404 | CL17 | HEK-293T | C299(0.77) | LDD1608 | [8] |
| LDCM0408 | CL20 | HEK-293T | C299(0.93) | LDD1612 | [8] |
| LDCM0409 | CL21 | HEK-293T | C299(0.90) | LDD1613 | [8] |
| LDCM0410 | CL22 | HEK-293T | C299(1.03); C119(0.75) | LDD1614 | [8] |
| LDCM0417 | CL29 | HEK-293T | C299(0.98) | LDD1621 | [8] |
| LDCM0421 | CL32 | HEK-293T | C299(1.07) | LDD1625 | [8] |
| LDCM0422 | CL33 | HEK-293T | C299(1.10) | LDD1626 | [8] |
| LDCM0423 | CL34 | HEK-293T | C299(0.95); C119(0.93) | LDD1627 | [8] |
| LDCM0431 | CL41 | HEK-293T | C299(0.91) | LDD1635 | [8] |
| LDCM0434 | CL44 | HEK-293T | C299(0.94) | LDD1638 | [8] |
| LDCM0435 | CL45 | HEK-293T | C299(0.99) | LDD1639 | [8] |
| LDCM0436 | CL46 | HEK-293T | C299(0.91); C119(0.71) | LDD1640 | [8] |
| LDCM0440 | CL5 | HEK-293T | C299(1.01) | LDD1644 | [8] |
| LDCM0444 | CL53 | HEK-293T | C299(0.95) | LDD1647 | [8] |
| LDCM0447 | CL56 | HEK-293T | C299(1.08) | LDD1650 | [8] |
| LDCM0448 | CL57 | HEK-293T | C299(1.18) | LDD1651 | [8] |
| LDCM0449 | CL58 | HEK-293T | C299(0.89); C119(0.90) | LDD1652 | [8] |
| LDCM0457 | CL65 | HEK-293T | C299(0.93) | LDD1660 | [8] |
| LDCM0460 | CL68 | HEK-293T | C299(1.05) | LDD1663 | [8] |
| LDCM0461 | CL69 | HEK-293T | C299(1.15) | LDD1664 | [8] |
| LDCM0463 | CL70 | HEK-293T | C299(0.99); C119(0.83) | LDD1666 | [8] |
| LDCM0470 | CL77 | HEK-293T | C299(1.00) | LDD1673 | [8] |
| LDCM0473 | CL8 | HEK-293T | C299(0.99) | LDD1676 | [8] |
| LDCM0474 | CL80 | HEK-293T | C299(0.96) | LDD1677 | [8] |
| LDCM0475 | CL81 | HEK-293T | C299(1.36) | LDD1678 | [8] |
| LDCM0476 | CL82 | HEK-293T | C299(0.87); C119(0.71) | LDD1679 | [8] |
| LDCM0483 | CL89 | HEK-293T | C299(0.90) | LDD1686 | [8] |
| LDCM0484 | CL9 | HEK-293T | C299(0.94) | LDD1687 | [8] |
| LDCM0487 | CL92 | HEK-293T | C299(0.92) | LDD1690 | [8] |
| LDCM0488 | CL93 | HEK-293T | C299(1.15) | LDD1691 | [8] |
| LDCM0489 | CL94 | HEK-293T | C299(1.06); C119(0.69) | LDD1692 | [8] |
| LDCM0107 | IAA | HeLa | H350(0.00); H116(0.00) | LDD0221 | [7] |
| LDCM0022 | KB02 | HEK-293T | C429(1.05) | LDD1492 | [8] |
| LDCM0023 | KB03 | HEK-293T | C429(0.92) | LDD1497 | [8] |
| LDCM0024 | KB05 | IGR37 | C429(2.42) | LDD3314 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [7] |
The Interaction Atlas With This Target
References
