Details of the Target
General Information of Target
| Target ID | LDTP11987 | |||||
|---|---|---|---|---|---|---|
| Target Name | Reticulophagy regulator 1 (RETREG1) | |||||
| Gene Name | RETREG1 | |||||
| Gene ID | 54463 | |||||
| Synonyms |
FAM134B; JK1; Reticulophagy regulator 1; Reticulophagy receptor 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAQKPKVDPHVGRLGYLQALVTEFQETQSQDAKEQVLANLANFAYDPSNYEYLRQLQVLD
LFLDSLSEENETLVEFAIGGLCNLCPDRANKEHILHAGGVPLIINCLSSPNEETVLSAIT TLMHLSPPGRSFLPELTATPVVQCMLRFSLSASARLRNLAQIFLEDFCSPRQVAEARSRQ AHSALGIPLPRSVAPRQR |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
RETREG family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane; Golgi apparatus, cis-Golgi network membrane
|
|||||
| Function |
Endoplasmic reticulum (ER)-anchored autophagy regulator which mediates ER delivery into lysosomes through sequestration into autophagosomes. Promotes membrane remodeling and ER scission via its membrane bending capacity and targets the fragments into autophagosomes via interaction with ATG8 family proteins. Active under basal conditions. Required for collagen quality control in a LIR motif-dependent manner. Required for long-term survival of nociceptive and autonomic ganglion neurons.; (Microbial infection) During SARS-CoV-2 infection, RETREG1-mediated reticulophagy is promoted by SARS-CoV-2 ORF3A protein. This induces endoplasmic reticulum stress and inflammatory responses and facilitates viral infection.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TG42 Probe Info |
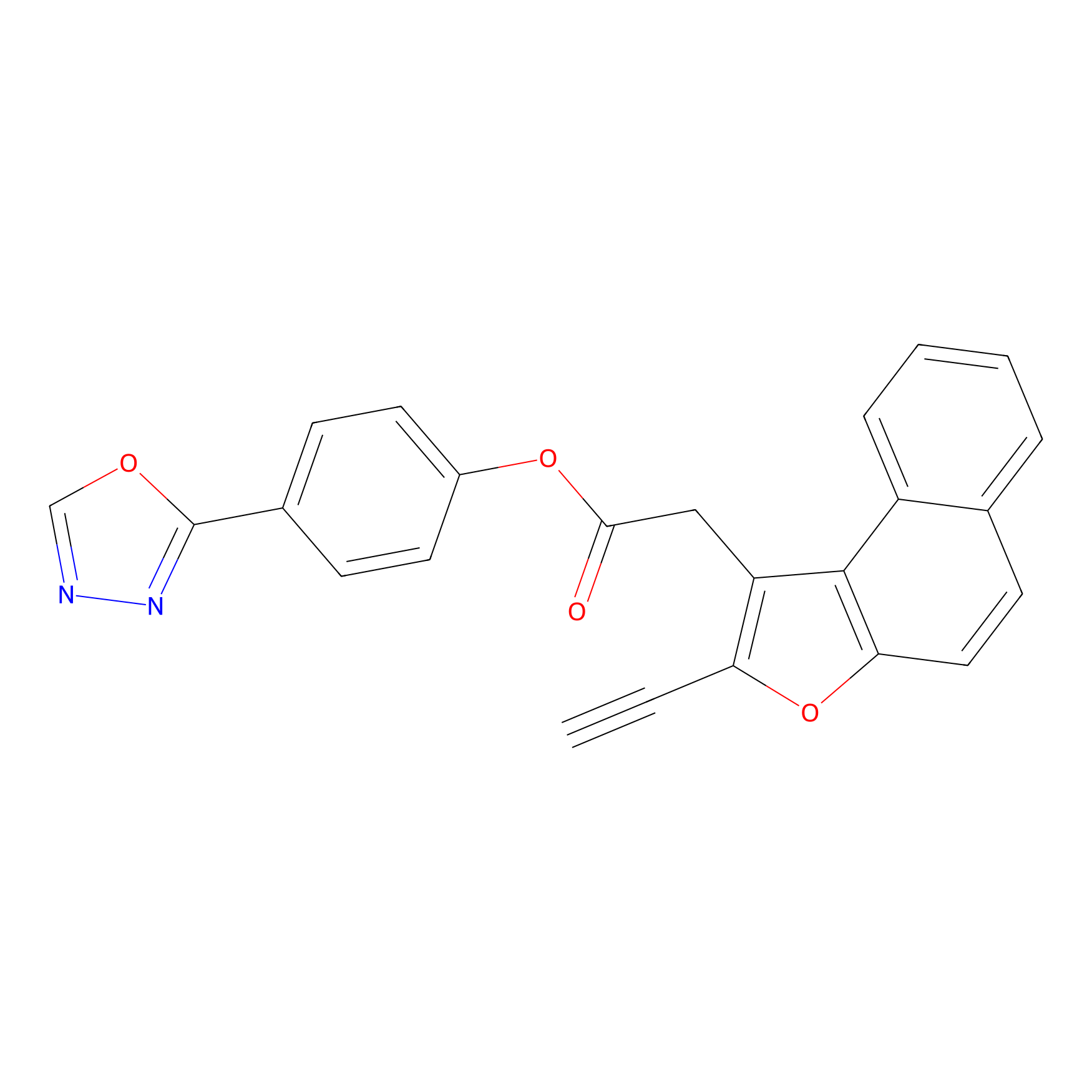 |
5.91 | LDD0043 | [1] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [2] | |
|
Johansson_61 Probe Info |
 |
_(20.00) | LDD1489 | [3] | |
|
NAIA_5 Probe Info |
 |
C289(1.01) | LDD2227 | [4] | |
|
AHL-Pu-1 Probe Info |
 |
C289(3.80) | LDD0168 | [5] | |
|
DBIA Probe Info |
 |
C235(1.02) | LDD1573 | [6] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0166 | [7] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [8] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C289(3.80) | LDD0168 | [5] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C289(2.22) | LDD0169 | [5] |
| LDCM0632 | CL-Sc | Hep-G2 | C289(1.01) | LDD2227 | [4] |
| LDCM0369 | CL100 | HEK-293T | C235(1.02) | LDD1573 | [6] |
| LDCM0373 | CL104 | HEK-293T | C235(1.03) | LDD1577 | [6] |
| LDCM0377 | CL108 | HEK-293T | C235(0.95) | LDD1581 | [6] |
| LDCM0382 | CL112 | HEK-293T | C235(1.01) | LDD1586 | [6] |
| LDCM0386 | CL116 | HEK-293T | C235(1.25) | LDD1590 | [6] |
| LDCM0391 | CL120 | HEK-293T | C235(0.99) | LDD1595 | [6] |
| LDCM0395 | CL124 | HEK-293T | C235(1.02) | LDD1599 | [6] |
| LDCM0399 | CL128 | HEK-293T | C235(0.99) | LDD1603 | [6] |
| LDCM0403 | CL16 | HEK-293T | C235(1.12) | LDD1607 | [6] |
| LDCM0416 | CL28 | HEK-293T | C235(1.01) | LDD1620 | [6] |
| LDCM0429 | CL4 | HEK-293T | C235(1.19) | LDD1633 | [6] |
| LDCM0430 | CL40 | HEK-293T | C235(1.04) | LDD1634 | [6] |
| LDCM0443 | CL52 | HEK-293T | C235(1.06) | LDD1646 | [6] |
| LDCM0456 | CL64 | HEK-293T | C235(1.31) | LDD1659 | [6] |
| LDCM0469 | CL76 | HEK-293T | C235(1.06) | LDD1672 | [6] |
| LDCM0482 | CL88 | HEK-293T | C235(1.11) | LDD1685 | [6] |
| LDCM0616 | Fragment61 | Jurkat | _(20.00) | LDD1489 | [3] |
The Interaction Atlas With This Target
References
