Details of the Target
General Information of Target
| Target ID | LDTP11804 | |||||
|---|---|---|---|---|---|---|
| Target Name | SH3 domain-binding glutamic acid-rich-like protein 3 (SH3BGRL3) | |||||
| Gene Name | SH3BGRL3 | |||||
| Gene ID | 83442 | |||||
| Synonyms |
SH3 domain-binding glutamic acid-rich-like protein 3; SH3 domain-binding protein 1; SH3BP-1; TNF inhibitory protein B1; TIP-B1 |
|||||
| 3D Structure | ||||||
| Sequence |
MGIWTSGTDIFLSLWEIYVSPRSPGWMDFIQHLGVCCLVALISVGLLSVAACWFLPSIIA
AAASWIITCVLLCCSKHARCFILLVFLSCGLREGRNALIAAGTGIVILGHVENIFHNFKG LLDGMTCNLRAKSFSIHFPLLKKYIEAIQWIYGLATPLSVFDDLVSWNQTLAVSLFSPSH VLEAQLNDSKGEVLSVLYQMATTTEVLSSLGQKLLAFAGLSLVLLGTGLFMKRFLGPCGW KYENIYITRQFVQFDERERHQQRPCVLPLNKEERRKYVIIPTFWPTPKERKNLGLFFLPI LIHLCIWVLFAAVDYLLYRLIFSVSKQFQSLPGFEVHLKLHGEKQGTQDIIHDSSFNISV FEPNCIPKPKFLLSETWVPLSVILLILVMLGLLSSILMQLKILVSASFYPSVERKRIQYL HAKLLKKRSKQPLGEVKRRLSLYLTKIHFWLPVLKMIRKKQMDMASADKS |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
SH3BGR family
|
|||||
| Subcellular location |
Cytoplasm, cytosol
|
|||||
| Function | Could act as a modulator of glutaredoxin biological activity (Probable). May play a role in cytoskeleton organization. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P1 Probe Info |
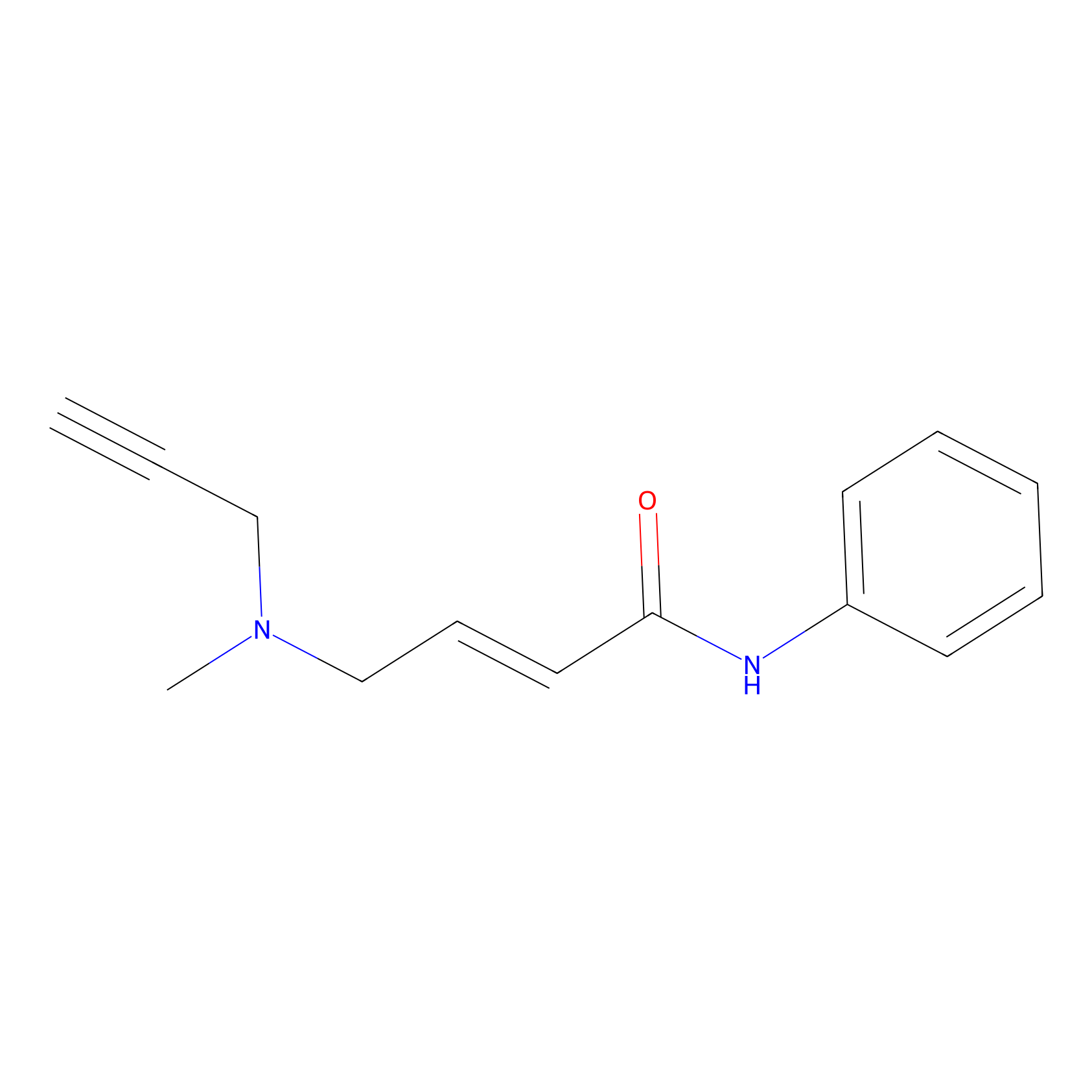 |
10.00 | LDD0448 | [1] | |
|
C-Sul Probe Info |
 |
4.55 | LDD0066 | [2] | |
|
FBP2 Probe Info |
 |
4.25 | LDD0323 | [3] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [4] | |
|
ONAyne Probe Info |
 |
K18(3.47) | LDD0274 | [5] | |
|
STPyne Probe Info |
 |
K18(8.62) | LDD0277 | [5] | |
|
AZ-9 Probe Info |
 |
E23(1.10); E49(0.43) | LDD2208 | [6] | |
|
DBIA Probe Info |
 |
C71(0.73) | LDD3345 | [7] | |
|
CY-1 Probe Info |
 |
D29(0.00); R26(0.00) | LDD0246 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0036 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [9] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0035 | [10] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [11] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [12] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [11] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
