Details of the Target
General Information of Target
| Target ID | LDTP10419 | |||||
|---|---|---|---|---|---|---|
| Target Name | Biorientation of chromosomes in cell division protein 1 (BOD1) | |||||
| Gene Name | BOD1 | |||||
| Gene ID | 91272 | |||||
| Synonyms |
FAM44B; Biorientation of chromosomes in cell division protein 1; Biorientation defective protein 1; Protein FAM44B |
|||||
| 3D Structure | ||||||
| Sequence |
MLKAVILIGGPQKGTRFRPLSFEVPKPLFPVAGVPMIQHHIEACAQVPGMQEILLIGFYQ
PDEPLTQFLEAAQQEFNLPVRYLQEFAPLGTGGGLYHFRDQILAGSPEAFFVLNADVCSD FPLSAMLEAHRRQRHPFLLLGTTANRTQSLNYGCIVENPQTHEVLHYVEKPSTFISDIIN CGIYLFSPEALKPLRDVFQRNQQDGQLEDSPGLWPGAGTIRLEQDVFSALAGQGQIYVHL TDGIWSQIKSAGSALYASRLYLSRYQDTHPERLAKHTPGGPWIRGNVYIHPTAKVAPSAV LGPNVSIGKGVTVGEGVRLRESIVLHGATLQEHTCVLHSIVGWGSTVGRWARVEGTPSDP NPNDPRARMDSESLFKDGKLLPAITILGCRVRIPAEVLILNSIVLPHKELSRSFTNQIIL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
BOD1 family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton, microtubule organizing center, centrosome
|
|||||
| Function | Required for proper chromosome biorientation through the detection or correction of syntelic attachments in mitotic spindles. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y82(10.38) | LDD0257 | [2] | |
|
STPyne Probe Info |
 |
K161(10.00) | LDD0277 | [3] | |
|
JZ128-DTB Probe Info |
 |
N.A. | LDD0462 | [4] | |
|
BTD Probe Info |
 |
C72(0.39) | LDD2091 | [5] | |
|
Curcusone 37 Probe Info |
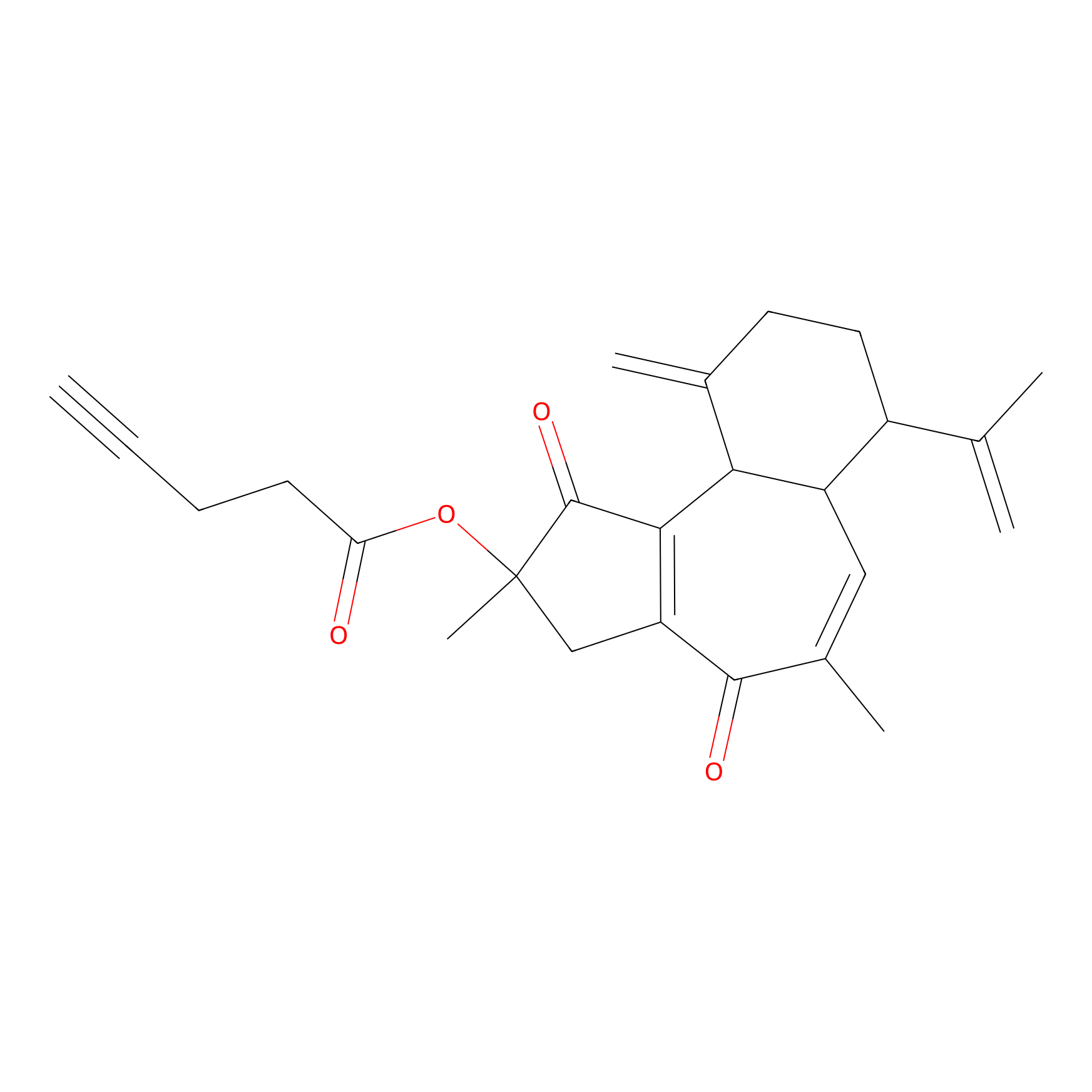 |
2.05 | LDD0188 | [6] | |
|
AHL-Pu-1 Probe Info |
 |
C72(2.66) | LDD0169 | [7] | |
|
HHS-465 Probe Info |
 |
Y82(3.69) | LDD2237 | [8] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [9] | |
|
WYneO Probe Info |
 |
N.A. | LDD0022 | [10] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [11] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | DM93 | C72(2.16) | LDD0170 | [7] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C72(2.66) | LDD0169 | [7] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C72(0.48) | LDD2113 | [5] |
| LDCM0156 | Aniline | NCI-H1299 | 15.00 | LDD0403 | [1] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C72(0.39) | LDD2091 | [5] |
| LDCM0033 | Curcusone1d | MCF-7 | 2.05 | LDD0188 | [6] |
| LDCM0179 | JZ128 | PC-3 | N.A. | LDD0462 | [4] |
| LDCM0509 | N-(4-bromo-3,5-dimethylphenyl)-2-nitroacetamide | MDA-MB-231 | C72(0.93) | LDD2102 | [5] |
| LDCM0499 | Nucleophilic fragment 12b | MDA-MB-231 | C72(0.76) | LDD2092 | [5] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C72(0.76) | LDD2096 | [5] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C72(0.75) | LDD2098 | [5] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C72(0.81) | LDD2101 | [5] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C72(0.74) | LDD2104 | [5] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C72(0.67) | LDD2108 | [5] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C72(0.99) | LDD2114 | [5] |
| LDCM0523 | Nucleophilic fragment 24b | MDA-MB-231 | C72(0.83) | LDD2116 | [5] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C72(1.54) | LDD2118 | [5] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C72(0.82) | LDD2120 | [5] |
| LDCM0529 | Nucleophilic fragment 27b | MDA-MB-231 | C72(0.89) | LDD2122 | [5] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C72(1.04) | LDD2124 | [5] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C72(0.69) | LDD2134 | [5] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C72(0.95) | LDD2135 | [5] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C72(0.63) | LDD2141 | [5] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C72(0.96) | LDD2146 | [5] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C72(0.43) | LDD2148 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| V-type proton ATPase subunit H (ATP6V1H) | V-ATPase H subunit family | Q9UI12 | |||
Other
References
