Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A2058 | SNV: p.H140Y | . | |||
| DANG | SNV: p.Y146D | DBIA Probe Info | |||
| HCT15 | Insertion: p.Y186VfsTer9 | DBIA Probe Info | |||
| JHH4 | SNV: p.K274R | DBIA Probe Info | |||
| MDAMB157 | SNV: p.C229R | DBIA Probe Info | |||
| NCIH446 | SNV: p.P41T | DBIA Probe Info | |||
| RPMI8226 | SNV: p.Q33L | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
FBPP2 Probe Info |
 |
5.10 | LDD0324 | [1] | |
|
N1 Probe Info |
 |
100.00 | LDD0242 | [2] | |
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [3] | |
|
STPyne Probe Info |
 |
K39(5.91) | LDD0277 | [4] | |
|
DBIA Probe Info |
 |
C82(1.03) | LDD3312 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [6] | |
|
IA-alkyne Probe Info |
 |
C31(0.00); C82(0.00) | LDD0162 | [7] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [8] | |
|
NAIA_4 Probe Info |
 |
C31(0.00); C82(0.00) | LDD2226 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Staurosporine capture compound Probe Info |
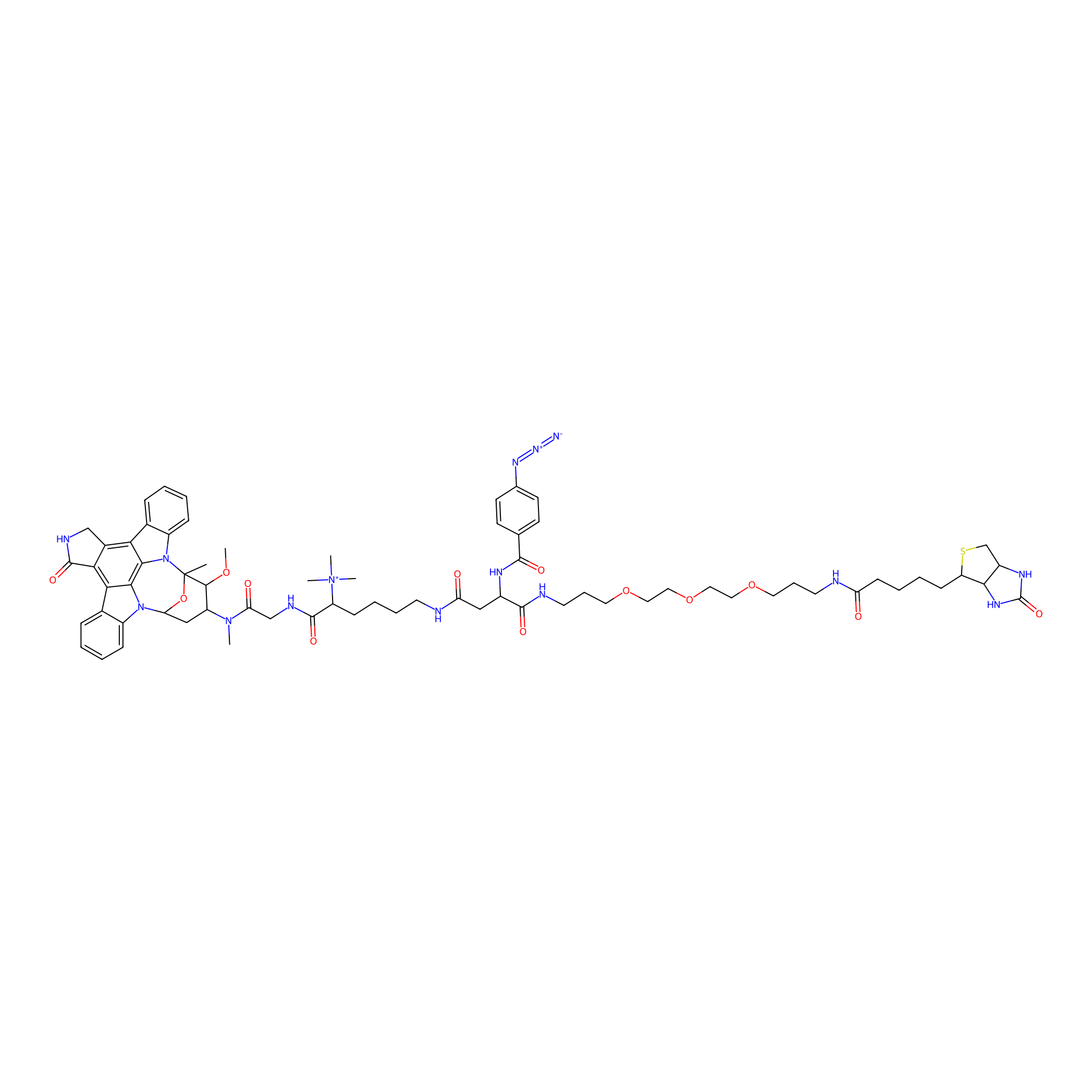 |
N.A. | LDD0083 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [6] |
| LDCM0632 | CL-Sc | Hep-G2 | C31(2.74) | LDD2227 | [9] |
| LDCM0022 | KB02 | 769-P | C82(0.81) | LDD2246 | [5] |
| LDCM0023 | KB03 | 8305C | C82(1.54) | LDD2665 | [5] |
| LDCM0024 | KB05 | HMCB | C82(1.03) | LDD3312 | [5] |
| LDCM0019 | Staurosporine | Hep-G2 | N.A. | LDD0083 | [10] |
The Interaction Atlas With This Target
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Amodiaquine | Small molecular drug | D04NQI | |||
| Diphenhydramine | Small molecular drug | D01FGR | |||
| Histamine | Small molecular drug | DB05381 | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Metoprine | Small molecular drug | D02AIM | |||
Investigative
References
