Details of the Target
General Information of Target
| Target ID | LDTP03781 | |||||
|---|---|---|---|---|---|---|
| Target Name | Transcription factor SOX-5 (SOX5) | |||||
| Gene Name | SOX5 | |||||
| Gene ID | 6660 | |||||
| Synonyms |
Transcription factor SOX-5 |
|||||
| 3D Structure | ||||||
| Sequence |
MLTDPDLPQEFERMSSKRPASPYGEADGEVAMVTSRQKVEEEESDGLPAFHLPLHVSFPN
KPHSEEFQPVSLLTQETCGHRTPTSQHNTMEVDGNKVMSSFAPHNSSTSPQKAEEGGRQS GESLSSTALGTPERRKGSLADVVDTLKQRKMEELIKNEPEETPSIEKLLSKDWKDKLLAM GSGNFGEIKGTPESLAEKERQLMGMINQLTSLREQLLAAHDEQKKLAASQIEKQRQQMEL AKQQQEQIARQQQQLLQQQHKINLLQQQIQVQGQLPPLMIPVFPPDQRTLAAAAQQGFLL PPGFSYKAGCSDPYPVQLIPTTMAAAAAATPGLGPLQLQQLYAAQLAAMQVSPGGKLPGI PQGNLGAAVSPTSIHTDKSTNSPPPKSKDEVAQPLNLSAKPKTSDGKSPTSPTSPHMPAL RINSGAGPLKASVPAALASPSARVSTIGYLNDHDAVTKAIQEARQMKEQLRREQQVLDGK VAVVNSLGLNNCRTEKEKTTLESLTQQLAVKQNEEGKFSHAMMDFNLSGDSDGSAGVSES RIYRESRGRGSNEPHIKRPMNAFMVWAKDERRKILQAFPDMHNSNISKILGSRWKAMTNL EKQPYYEEQARLSKQHLEKYPDYKYKPRPKRTCLVDGKKLRIGEYKAIMRNRRQEMRQYF NVGQQAQIPIATAGVVYPGAIAMAGMPSPHLPSEHSSVSSSPEPGMPVIQSTYGVKGEEP HIKEEIQAEDINGEIYDEYDEEEDDPDVDYGSDSENHIAGQAN |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Transcription factor
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Transcription factor involved in chondrocytes differentiation and cartilage formation. Specifically binds the 5'-AACAAT-3' DNA motif present in enhancers and super-enhancers and promotes expression of genes important for chondrogenesis, including cartilage matrix protein-coding genes, such as COL2A1 and AGC1. Required for overt chondrogenesis when condensed prechondrocytes differentiate into early stage chondrocytes: SOX5 and SOX6 cooperatively bind with SOX9 on active enhancers and super-enhancers associated with cartilage-specific genes, and thereby potentiate SOX9's ability to transactivate. Not involved in precartilaginous condensation, the first step in chondrogenesis, during which skeletal progenitors differentiate into prechondrocytes. Together with SOX6, required to form and maintain a pool of highly proliferating chondroblasts between epiphyses and metaphyses, to form columnar chondroblasts, delay chondrocyte prehypertrophy but promote hypertrophy, and to delay terminal differentiation of chondrocytes on contact with ossification fronts. Binds to the proximal promoter region of the myelin protein MPZ gene.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A549 | SNV: p.E703D | . | |||
| CAL148 | SNV: p.E114K | . | |||
| HT115 | SNV: p.R288W | . | |||
| KYSE150 | SNV: p.H220Y | . | |||
| LN229 | SNV: p.E755Q | . | |||
| LNCaP clone FGC | SNV: p.D377Y | . | |||
| MOLT4 | SNV: p.R650H | . | |||
| NCIH446 | SNV: p.A577V | . | |||
| SNU1 | SNV: p.V635L | DBIA Probe Info | |||
| SUIT2 | SNV: p.P579S | . | |||
| SW1271 | SNV: p.T321N | . | |||
| TYKNU | SNV: p.H453N; p.R541S | . | |||
| VCAP | SNV: p.M686T | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
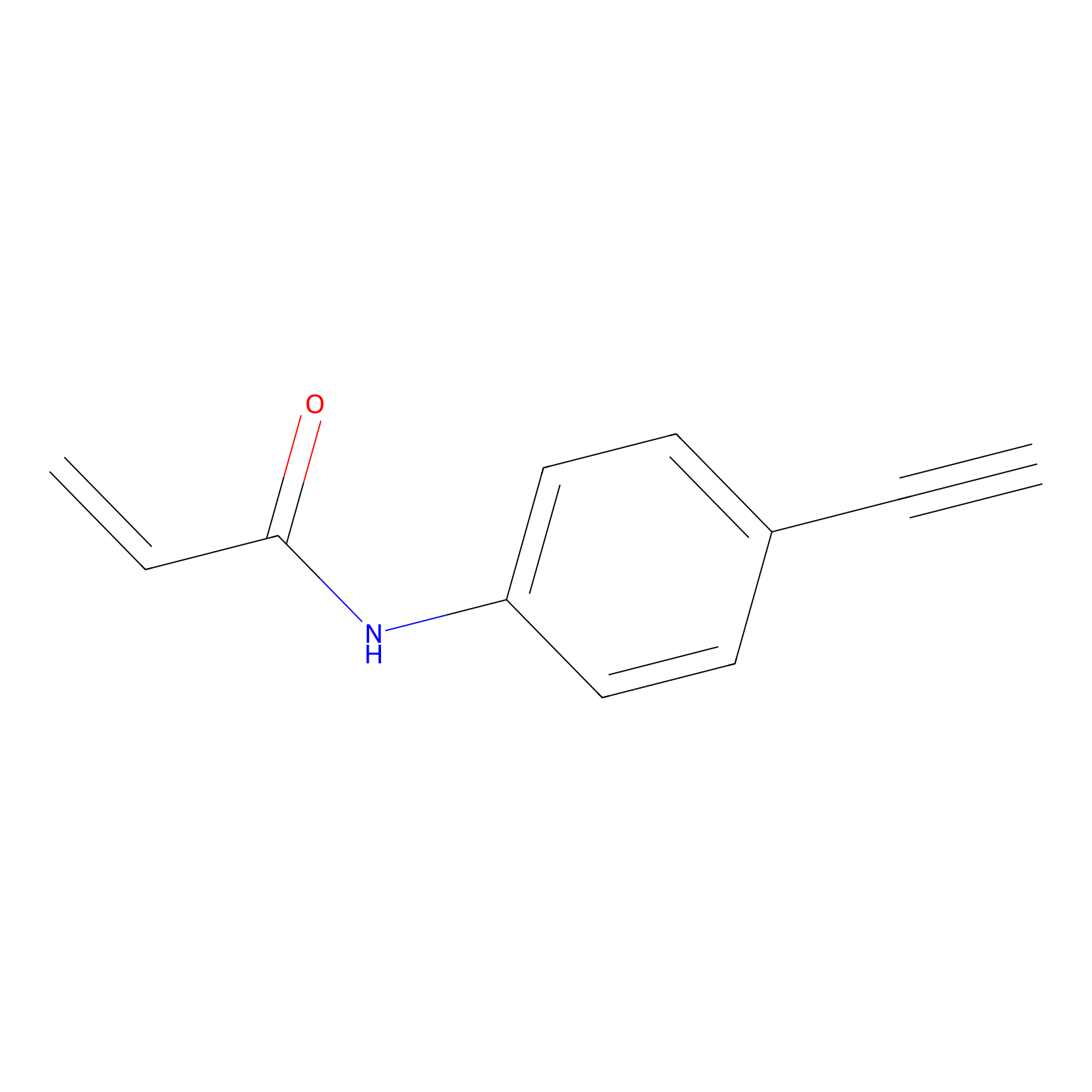 |
10.00 | LDD0449 | [1] | |
|
DBIA Probe Info |
 |
C492(2.72) | LDD3332 | [2] | |
|
EA-probe Probe Info |
 |
N.A. | LDD0440 | [3] | |
|
IA-alkyne Probe Info |
 |
C492(0.68) | LDD2183 | [4] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [5] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0175 | Ethacrynic acid | HeLa | N.A. | LDD0440 | [3] |
| LDCM0625 | F8 | Ramos | C492(2.38) | LDD2187 | [4] |
| LDCM0572 | Fragment10 | Ramos | C492(0.66) | LDD2189 | [4] |
| LDCM0574 | Fragment12 | Ramos | C492(0.73) | LDD2191 | [4] |
| LDCM0575 | Fragment13 | Ramos | C492(0.85) | LDD2192 | [4] |
| LDCM0576 | Fragment14 | Ramos | C492(1.23) | LDD2193 | [4] |
| LDCM0580 | Fragment21 | Ramos | C492(0.66) | LDD2195 | [4] |
| LDCM0582 | Fragment23 | Ramos | C492(0.39) | LDD2196 | [4] |
| LDCM0578 | Fragment27 | Ramos | C492(0.87) | LDD2197 | [4] |
| LDCM0586 | Fragment28 | Ramos | C492(0.52) | LDD2198 | [4] |
| LDCM0588 | Fragment30 | Ramos | C492(0.91) | LDD2199 | [4] |
| LDCM0589 | Fragment31 | Ramos | C492(0.78) | LDD2200 | [4] |
| LDCM0590 | Fragment32 | Ramos | C492(0.95) | LDD2201 | [4] |
| LDCM0468 | Fragment33 | Ramos | C492(1.02) | LDD2202 | [4] |
| LDCM0596 | Fragment38 | Ramos | C492(0.58) | LDD2203 | [4] |
| LDCM0566 | Fragment4 | Ramos | C492(1.07) | LDD2184 | [4] |
| LDCM0610 | Fragment52 | Ramos | C492(0.61) | LDD2204 | [4] |
| LDCM0614 | Fragment56 | Ramos | C492(0.57) | LDD2205 | [4] |
| LDCM0569 | Fragment7 | Ramos | C492(0.92) | LDD2186 | [4] |
| LDCM0571 | Fragment9 | Ramos | C492(0.82) | LDD2188 | [4] |
| LDCM0022 | KB02 | A101D | C492(2.34) | LDD2250 | [2] |
| LDCM0023 | KB03 | Ramos | C492(0.68) | LDD2183 | [4] |
| LDCM0024 | KB05 | MKN45 | C492(2.72) | LDD3332 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
Other
References
