Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DA-P2 Probe Info |
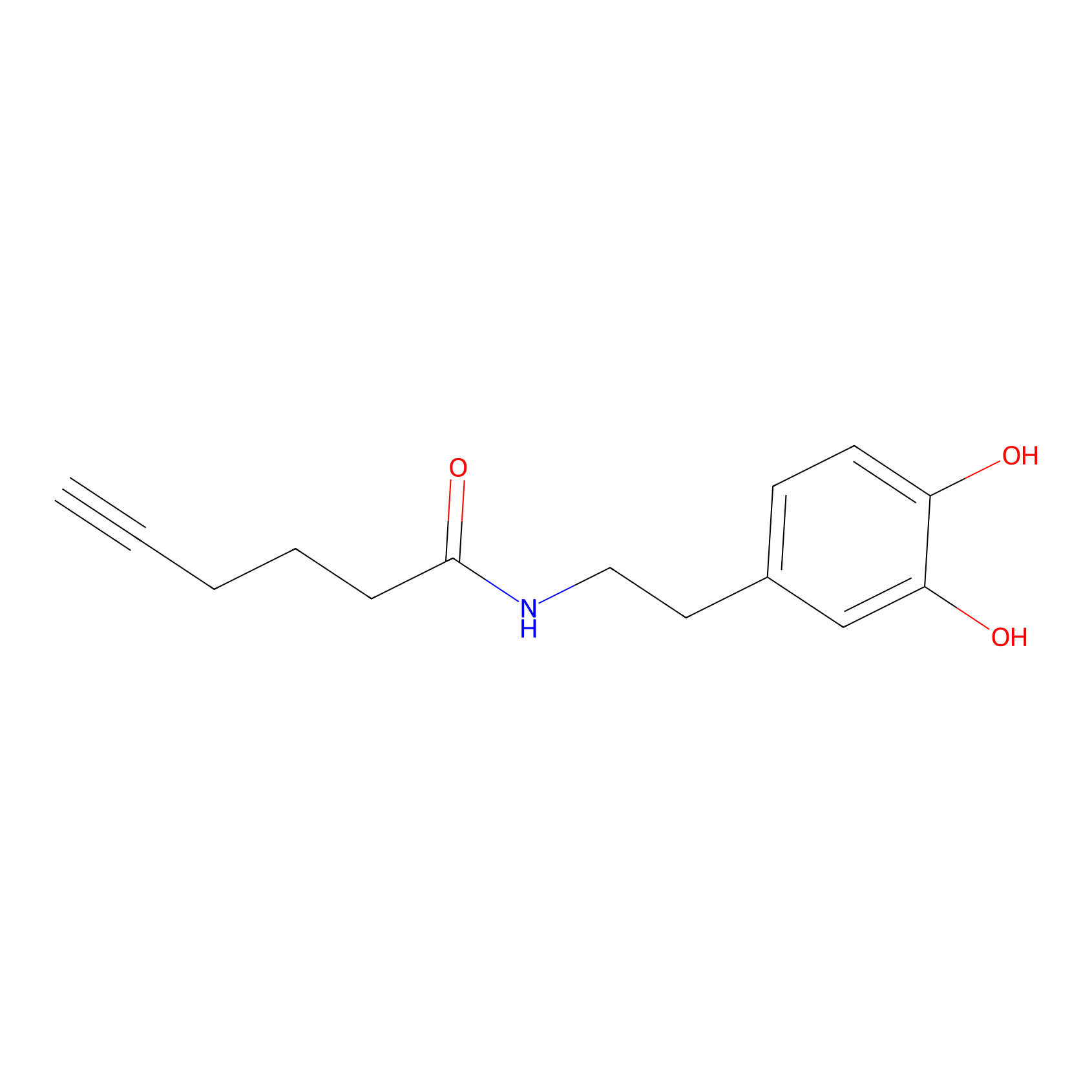 |
13.00 | LDD0348 | [1] | |
|
HRP Probe Info |
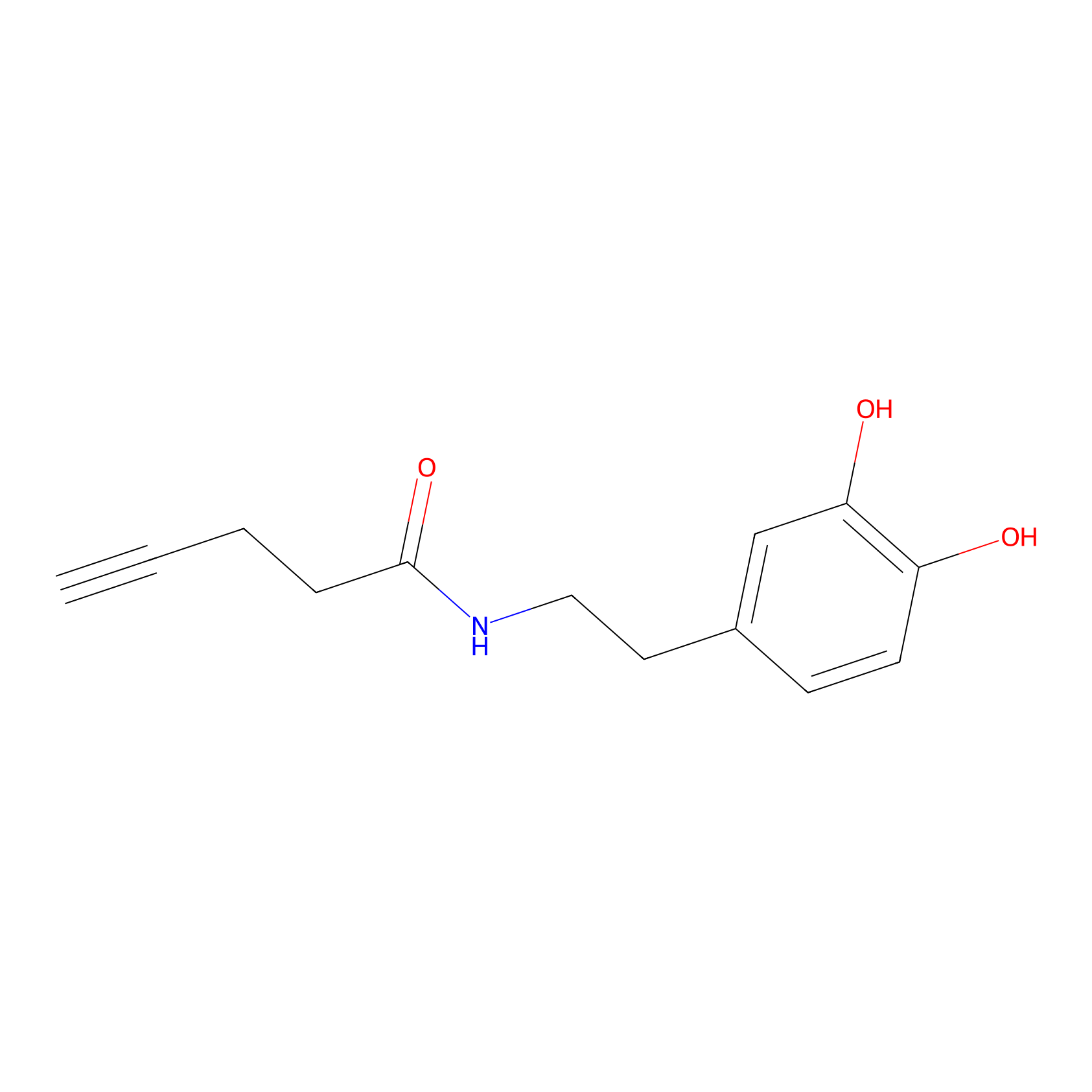 |
6.46 | LDD0347 | [1] | |
|
DA-P3 Probe Info |
 |
6.16 | LDD0185 | [1] | |
|
IPM Probe Info |
 |
N.A. | LDD0147 | [2] | |
|
AOyne Probe Info |
 |
9.30 | LDD0443 | [3] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Endoplasmic reticulum chaperone BiP (HSPA5) | Heat shock protein 70 family | P11021 | |||
Transcription factor
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| CREB-regulated transcription coactivator 2 (CRTC2) | TORC family | Q53ET0 | |||
References
