Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K125(1.34) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K240(10.00) | LDD0277 | [2] | |
|
AZ-9 Probe Info |
 |
E413(0.78); E114(1.05); D117(1.11); E387(10.00) | LDD2208 | [3] | |
|
DBIA Probe Info |
 |
C333(1.82) | LDD3401 | [4] | |
|
HHS-465 Probe Info |
 |
Y388(0.00); K407(0.00); K109(0.00) | LDD2240 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C238 Probe Info |
 |
17.15 | LDD1911 | [6] | |
|
C239 Probe Info |
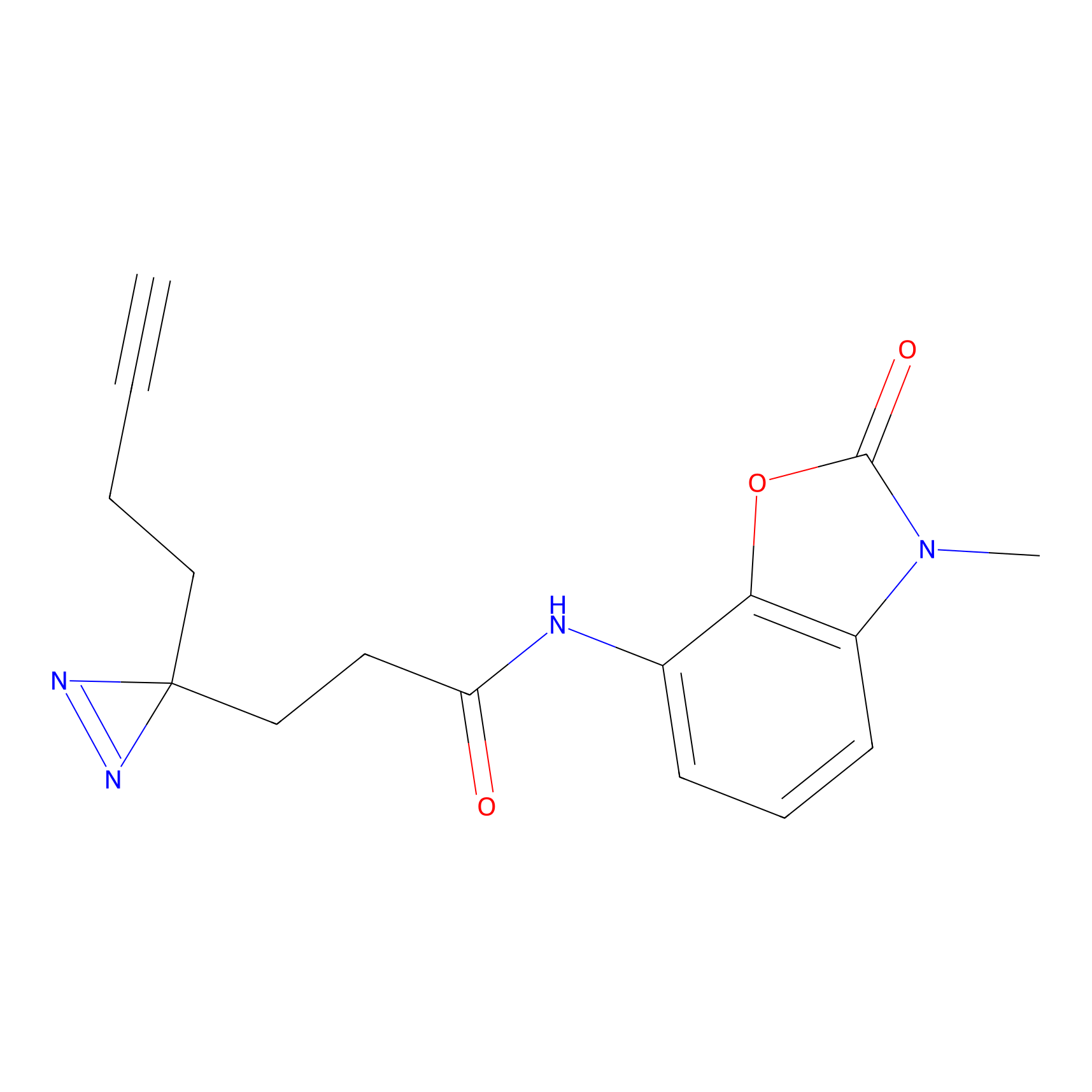 |
5.62 | LDD1912 | [6] | |
|
C390 Probe Info |
 |
28.44 | LDD2049 | [6] | |
|
C391 Probe Info |
 |
43.41 | LDD2050 | [6] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor LBX1 (LBX1) | . | P52954 | |||
Other
References
