Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P11 Probe Info |
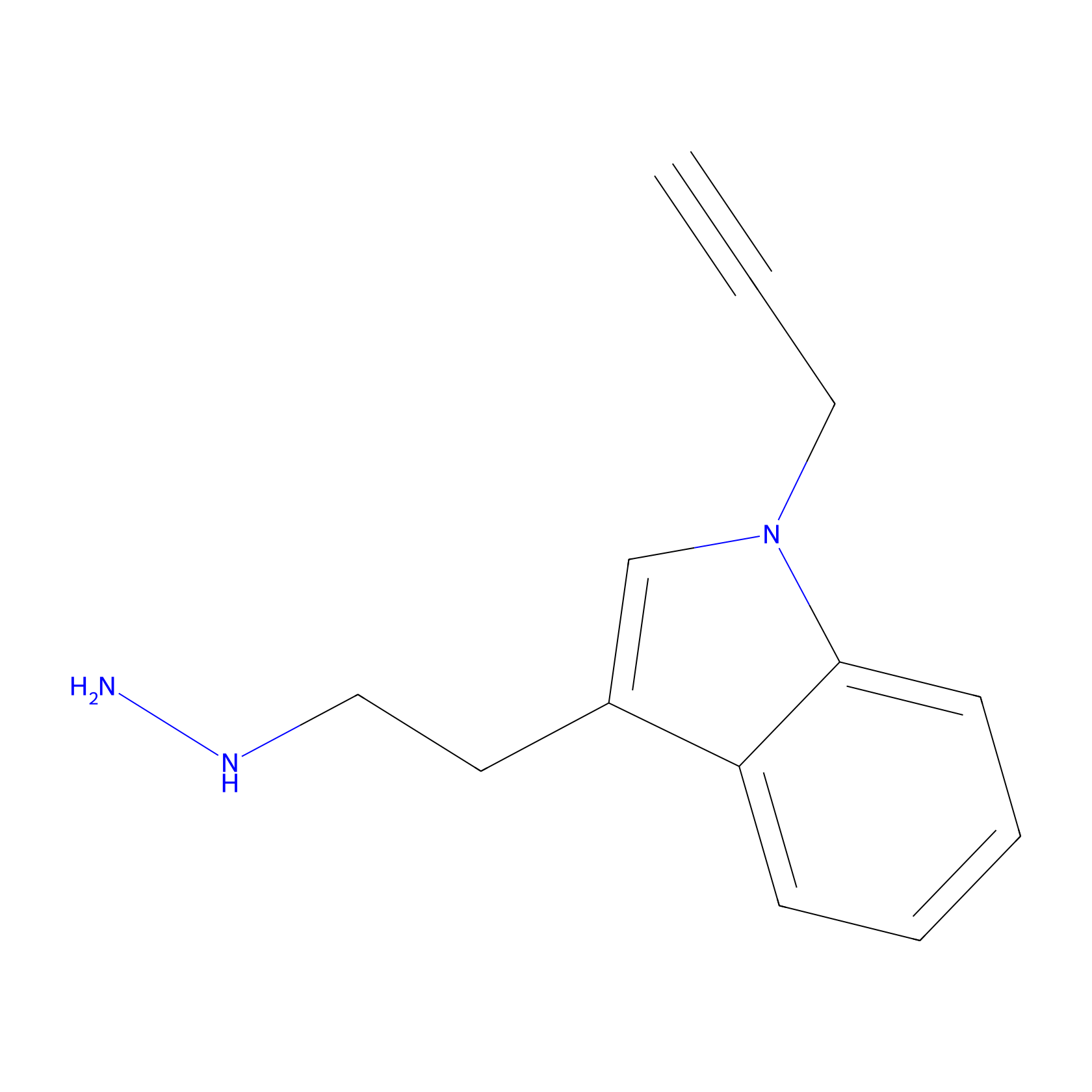 |
20.00 | LDD0201 | [1] | |
|
DBIA Probe Info |
 |
C272(1.48) | LDD3330 | [2] | |
|
FBPP2 Probe Info |
 |
5.70 | LDD0054 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C172(3.87) | LDD0171 | [4] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [5] | |
|
IA-alkyne Probe Info |
 |
C272(0.00); C172(0.00) | LDD0162 | [6] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [7] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | DM93 | C172(3.87) | LDD0171 | [4] |
| LDCM0281 | AC21 | HEK-293T | C141(0.77) | LDD1520 | [9] |
| LDCM0289 | AC29 | HEK-293T | C141(1.02) | LDD1528 | [9] |
| LDCM0298 | AC37 | HEK-293T | C141(1.40) | LDD1537 | [9] |
| LDCM0307 | AC45 | HEK-293T | C141(1.06) | LDD1546 | [9] |
| LDCM0312 | AC5 | HEK-293T | C141(1.00) | LDD1551 | [9] |
| LDCM0316 | AC53 | HEK-293T | C141(0.89) | LDD1555 | [9] |
| LDCM0325 | AC61 | HEK-293T | C141(0.78) | LDD1564 | [9] |
| LDCM0248 | AKOS034007472 | HEK-293T | C141(1.00) | LDD1511 | [9] |
| LDCM0088 | C45 | HEK-293T | 20.00 | LDD0201 | [1] |
| LDCM0367 | CL1 | HEK-293T | C102(1.06) | LDD1571 | [9] |
| LDCM0370 | CL101 | HEK-293T | C102(0.90) | LDD1574 | [9] |
| LDCM0374 | CL105 | HEK-293T | C102(0.78) | LDD1578 | [9] |
| LDCM0378 | CL109 | HEK-293T | C102(0.68) | LDD1582 | [9] |
| LDCM0383 | CL113 | HEK-293T | C102(0.89) | LDD1587 | [9] |
| LDCM0387 | CL117 | HEK-293T | C102(0.83) | LDD1591 | [9] |
| LDCM0392 | CL121 | HEK-293T | C102(0.96) | LDD1596 | [9] |
| LDCM0396 | CL125 | HEK-293T | C102(0.99) | LDD1600 | [9] |
| LDCM0400 | CL13 | HEK-293T | C102(0.95) | LDD1604 | [9] |
| LDCM0409 | CL21 | HEK-293T | C141(0.71) | LDD1613 | [9] |
| LDCM0413 | CL25 | HEK-293T | C102(0.94) | LDD1617 | [9] |
| LDCM0422 | CL33 | HEK-293T | C141(1.18) | LDD1626 | [9] |
| LDCM0426 | CL37 | HEK-293T | C102(1.03) | LDD1630 | [9] |
| LDCM0435 | CL45 | HEK-293T | C141(0.78) | LDD1639 | [9] |
| LDCM0439 | CL49 | HEK-293T | C102(0.82) | LDD1643 | [9] |
| LDCM0448 | CL57 | HEK-293T | C141(1.15) | LDD1651 | [9] |
| LDCM0453 | CL61 | HEK-293T | C102(0.83) | LDD1656 | [9] |
| LDCM0461 | CL69 | HEK-293T | C141(1.04) | LDD1664 | [9] |
| LDCM0466 | CL73 | HEK-293T | C102(0.93) | LDD1669 | [9] |
| LDCM0475 | CL81 | HEK-293T | C141(1.05) | LDD1678 | [9] |
| LDCM0479 | CL85 | HEK-293T | C102(1.13) | LDD1682 | [9] |
| LDCM0484 | CL9 | HEK-293T | C141(0.80) | LDD1687 | [9] |
| LDCM0488 | CL93 | HEK-293T | C141(1.34) | LDD1691 | [9] |
| LDCM0492 | CL97 | HEK-293T | C102(0.94) | LDD1695 | [9] |
| LDCM0022 | KB02 | KP-4 | C272(1.71) | LDD2428 | [2] |
| LDCM0023 | KB03 | MIA PaCa-2 | C272(2.06) | LDD2913 | [2] |
| LDCM0024 | KB05 | MIA PaCa-2 | C272(1.48) | LDD3330 | [2] |
| LDCM0008 | Tranylcypromine | SH-SY5Y | 5.70 | LDD0054 | [3] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Kallikrein-6 (KLK6) | Peptidase S1 family | Q92876 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Cytoplasmic dynein 1 heavy chain 1 (DYNC1H1) | Dynein heavy chain family | Q14204 | |||
The Drug(s) Related To This Target
Investigative
References
