Details of the Target
General Information of Target
| Target ID | LDTP00119 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tubulin beta 8B (TUBB8B) | |||||
| Gene Name | TUBB8B | |||||
| Synonyms |
Tubulin beta 8B |
|||||
| 3D Structure | ||||||
| Sequence |
MREIVLTQTGQCGNQIGAKFWEVISDEHAIDSAGTYHGDSHLQLERINVHHHEASGGRYV
PRAVLVDLEPGTMDSVHSGPFGQVFRPDNFISGQCGAGNNWAKGRYTEGAELTESVMDVV RKEAESCDCLQGFQLTHSLGGGTGSGMGTLLISKIREEYPDRIINTFSILPSPKVSDTVV EPYNATLSVHQLIENADETFCIDNEALYDICSRTLKLPTPTYGDLNHLVSATMSGVTTCL RFPGQLNADLRKLAVNMVPFPRLHFFMPGFAPLTSRGSQQYRALTVAELTQQMFDAKNMM AACDPRHGCYLTVAAIFRGRMPMREVDEQMFNIQDKNSSYFADWFPDNVKTAVCDIPPRG LKMSATFIGNNAAIQELFTCVSEQFTAMFRRKAFLHWYTGEGMDEMEFTEAESNMNDLVS EYQQYQDATAEEEEDEEYAEEEVA |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Tubulin family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton
|
|||||
| Function |
Tubulin is the major constituent of microtubules, a cylinder consisting of laterally associated linear protofilaments composed of alpha- and beta-tubulin heterodimers. Microtubules grow by the addition of GTP-tubulin dimers to the microtubule end, where a stabilizing cap forms. Below the cap, tubulin dimers are in GDP-bound state, owing to GTPase activity of alpha-tubulin.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
OPA-S-S-alkyne Probe Info |
 |
K252(2.04) | LDD3494 | [1] | |
|
JZ128-DTB Probe Info |
 |
C129(0.00); C127(0.00) | LDD0462 | [2] | |
|
THZ1-DTB Probe Info |
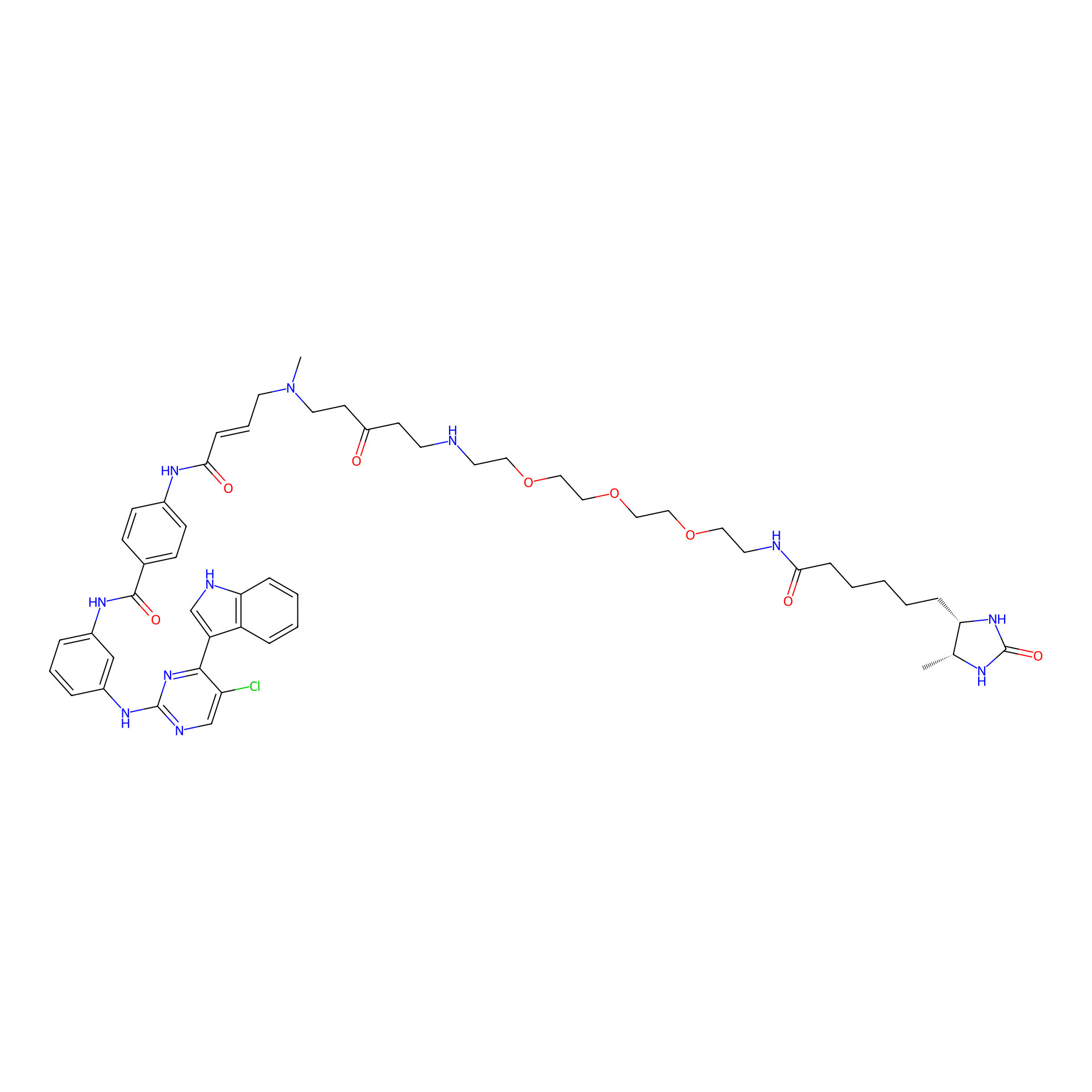 |
C303(1.20); C127(1.08); C354(1.05); C129(1.02) | LDD0460 | [2] | |
|
BTD Probe Info |
 |
C12(0.85) | LDD2094 | [3] | |
|
AHL-Pu-1 Probe Info |
 |
C129(2.47); C303(2.15) | LDD0168 | [4] | |
|
HHS-475 Probe Info |
 |
Y281(0.79) | LDD0264 | [5] | |
|
HHS-465 Probe Info |
 |
Y159(8.69); Y222(6.05); Y281(7.61) | LDD2237 | [6] | |
|
DBIA Probe Info |
 |
C127(1.03); C129(1.15); C127(0.98); C129(0.98) | LDD1492 | [7] | |
|
5E-2FA Probe Info |
 |
H137(0.00); H264(0.00); H227(0.00) | LDD2235 | [8] | |
|
m-APA Probe Info |
 |
H137(0.00); H264(0.00); H227(0.00) | LDD2231 | [8] | |
|
IA-alkyne Probe Info |
 |
C239(0.00); C354(0.00); C303(0.00); C127(0.00) | LDD0166 | [9] | |
|
JW-RF-010 Probe Info |
 |
C303(0.00); C129(0.00); C354(0.00); C127(0.00) | LDD0026 | [10] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [11] | |
|
ENE Probe Info |
 |
C303(0.00); C354(0.00) | LDD0006 | [12] | |
|
IPM Probe Info |
 |
C303(0.00); C354(0.00); C127(0.00); C129(0.00) | LDD0147 | [10] | |
|
VSF Probe Info |
 |
C129(0.00); C127(0.00) | LDD0007 | [12] | |
|
NAIA_5 Probe Info |
 |
C127(0.00); C239(0.00); C12(0.00); C303(0.00) | LDD2223 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C129(2.47); C303(2.15) | LDD0168 | [4] |
| LDCM0630 | CCW28-3 | 231MFP | C239(1.48) | LDD2214 | [13] |
| LDCM0116 | HHS-0101 | DM93 | Y281(0.79) | LDD0264 | [5] |
| LDCM0117 | HHS-0201 | DM93 | Y281(0.80) | LDD0265 | [5] |
| LDCM0118 | HHS-0301 | DM93 | Y281(0.85) | LDD0266 | [5] |
| LDCM0119 | HHS-0401 | DM93 | Y281(0.91) | LDD0267 | [5] |
| LDCM0120 | HHS-0701 | DM93 | Y281(0.76) | LDD0268 | [5] |
| LDCM0179 | JZ128 | PC-3 | C129(0.00); C127(0.00) | LDD0462 | [2] |
| LDCM0022 | KB02 | HEK-293T | C127(1.03); C129(1.15); C127(0.98); C129(0.98) | LDD1492 | [7] |
| LDCM0023 | KB03 | HEK-293T | C127(0.99); C129(1.13); C127(0.94); C129(0.94) | LDD1497 | [7] |
| LDCM0024 | KB05 | HEK-293T | C127(1.12); C129(1.13); C127(1.04); C129(1.04) | LDD1502 | [7] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C12(0.85) | LDD2094 | [3] |
| LDCM0505 | Nucleophilic fragment 15b | MDA-MB-231 | C12(1.17) | LDD2098 | [3] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C129(1.13) | LDD2206 | [14] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C12(15.00); C127(0.48); C129(0.48); C127(0.20) | LDD2207 | [14] |
| LDCM0021 | THZ1 | HeLa S3 | C303(1.20); C127(1.08); C354(1.05); C129(1.02) | LDD0460 | [2] |
References
