Details of the Target
General Information of Target
| Target ID | LDTP13920 | |||||
|---|---|---|---|---|---|---|
| Target Name | WW domain-binding protein 11 (WBP11) | |||||
| Gene Name | WBP11 | |||||
| Gene ID | 51729 | |||||
| Synonyms |
NPWBP; SIPP1; SNP70; WW domain-binding protein 11; WBP-11; Npw38-binding protein; NpwBP; SH3 domain-binding protein SNP70; Splicing factor that interacts with PQBP-1 and PP1 |
|||||
| 3D Structure | ||||||
| Sequence |
MSKTNKSKSGSRSSRSRSASRSRSRSFSKSRSRSRSLSRSRKRRLSSRSRSRSYSPAHNR
ERNHPRVYQNRDFRGHNRGYRRPYYFRGRNRGFYPWGQYNRGGYGNYRSNWQNYRQAYSP RRGRSRSRSPKRRSPSPRSRSHSRNSDKSSSDRSRRSSSSRSSSNHSRVESSKRKSAKEK KSSSKDSRPSQAAGDNQGDEAKEQTFSGGTSQDTKASESSKPWPDATYGTGSASRASAVS ELSPRERSPALKSPLQSVVVRRRSPRPSPVPKPSPPLSSTSQMGSTLPSGAGYQSGTHQG QFDHGSGSLSPSKKSPVGKSPPSTGSTYGSSQKEESAASGGAAYTKRYLEEQKTENGKDK EQKQTNTDKEKIKEKGSFSDTGLGDGKMKSDSFAPKTDSEKPFRGSQSPKRYKLRDDFEK KMADFHKEEMDDQDKDKAKGRKESEFDDEPKFMSKVIGANKNQEEEKSGKWEGLVYAPPG KEKQRKTEELEEESFPERSKKEDRGKRSEGGHRGFVPEKNFRVTAYKAVQEKSSSPPPRK TSESRDKLGAKGDFPTGKSSFSITREAQVNVRMDSFDEDLARPSGLLAQERKLCRDLVHS NKKEQEFRSIFQHIQSAQSQRSPSELFAQHIVTIVHHVKEHHFGSSGMTLHERFTKYLKR GTEQEAAKNKKSPEIHRRIDISPSTFRKHGLAHDEMKSPREPGYKAEGKYKDDPVDLRLD IERRKKHKERDLKRGKSRESVDSRDSSHSRERSAEKTEKTHKGSKKQKKHRRARDRSRSS SSSSQSSHSYKAEEYTEETEEREESTTGFDKSRLGTKDFVGPSERGGGRARGTFQFRARG RGWGRGNYSGNNNNNSNNDFQKRNREEEWDPEYTPKSKKYYLHDDREGEGSDKWVSRGRG RGAFPRGRGRFMFRKSSTSPKWAHDKFSGEEGEIEDDESGTENREEKDNIQPTTE |
|||||
| Target Bioclass |
Other
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function | Activates pre-mRNA splicing. May inhibit PP1 phosphatase activity. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AZ-9 Probe Info |
 |
10.00 | LDD0393 | [1] | |
|
STPyne Probe Info |
 |
K169(6.62); K81(6.64); K98(5.00) | LDD0277 | [2] | |
|
Probe 1 Probe Info |
 |
Y124(16.85); Y173(33.18); Y236(5.60) | LDD3495 | [3] | |
|
Alkylaryl probe 3 Probe Info |
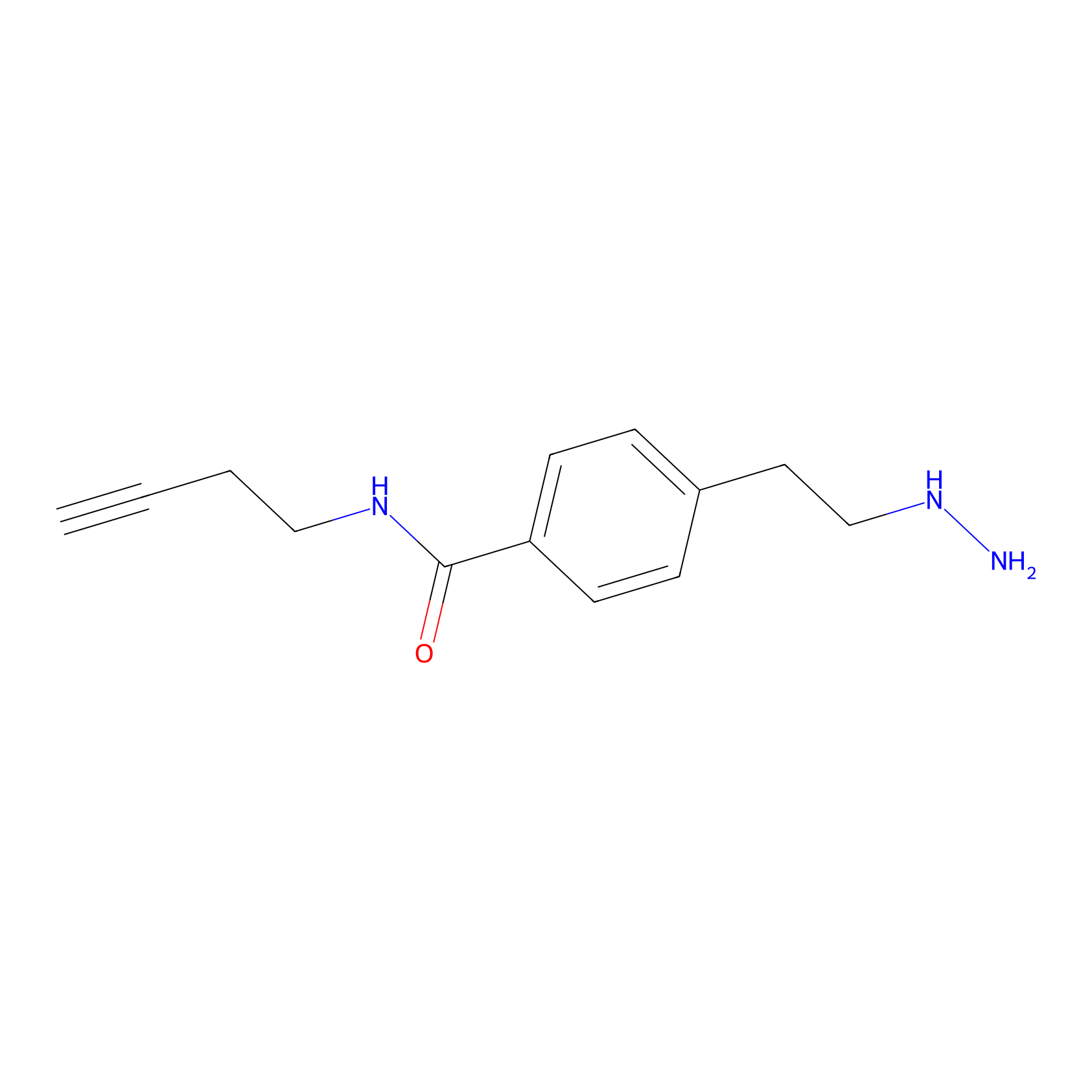 |
20.00 | LDD0382 | [4] | |
|
HHS-475 Probe Info |
 |
Y173(0.84) | LDD0264 | [5] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [6] | |
|
5E-2FA Probe Info |
 |
H188(0.00); H133(0.00) | LDD2235 | [7] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [8] | |
|
m-APA Probe Info |
 |
H188(0.00); H133(0.00); H146(0.00) | LDD2231 | [7] | |
|
NHS Probe Info |
 |
N.A. | LDD0010 | [9] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [10] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [11] | |
|
HHS-482 Probe Info |
 |
Y173(0.78) | LDD2239 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [6] |
| LDCM0116 | HHS-0101 | DM93 | Y173(0.84) | LDD0264 | [5] |
| LDCM0117 | HHS-0201 | DM93 | Y173(0.95) | LDD0265 | [5] |
| LDCM0118 | HHS-0301 | DM93 | Y173(1.30) | LDD0266 | [5] |
| LDCM0119 | HHS-0401 | DM93 | Y173(1.15) | LDD0267 | [5] |
| LDCM0120 | HHS-0701 | DM93 | Y173(1.10) | LDD0268 | [5] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [6] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [6] |
| LDCM0099 | Phenelzine | HEK-293T | 20.00 | LDD0382 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Other
References
