Details of the Target
General Information of Target
| Target ID | LDTP09651 | |||||
|---|---|---|---|---|---|---|
| Target Name | Choline transporter-like protein 1 (SLC44A1) | |||||
| Gene Name | SLC44A1 | |||||
| Gene ID | 23446 | |||||
| Synonyms |
CD92; CDW92; CTL1; Choline transporter-like protein 1; CDw92; Solute carrier family 44 member 1; CD antigen CD92 |
|||||
| 3D Structure | ||||||
| Sequence |
MGCCSSASSAAQSSKREWKPLEDRSCTDIPWLLLFILFCIGMGFICGFSIATGAAARLVS
GYDSYGNICGQKNTKLEAIPNSGMDHTQRKYVFFLDPCNLDLINRKIKSVALCVAACPRQ ELKTLSDVQKFAEINGSALCSYNLKPSEYTTSPKSSVLCPKLPVPASAPIPFFHRCAPVN ISCYAKFAEALITFVSDNSVLHRLISGVMTSKEIILGLCLLSLVLSMILMVIIRYISRVL VWILTILVILGSLGGTGVLWWLYAKQRRSPKETVTPEQLQIAEDNLRALLIYAISATVFT VILFLIMLVMRKRVALTIALFHVAGKVFIHLPLLVFQPFWTFFALVLFWVYWIMTLLFLG TTGSPVQNEQGFVEFKISGPLQYMWWYHVVGLIWISEFILACQQMTVAGAVVTYYFTRDK RNLPFTPILASVNRLIRYHLGTVAKGSFIITLVKIPRMILMYIHSQLKGKENACARCVLK SCICCLWCLEKCLNYLNQNAYTATAINSTNFCTSAKDAFVILVENALRVATINTVGDFML FLGKVLIVCSTGLAGIMLLNYQQDYTVWVLPLIIVCLFAFLVAHCFLSIYEMVVDVLFLC FAIDTKYNDGSPGREFYMDKVLMEFVENSRKAMKEAGKGGVADSRELKPMASGASSA |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
CTL (choline transporter-like) family
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
Choline/H+ antiporter. Also acts as a high-affinity ethanolamine/H+ antiporter, regulating the supply of extracellular ethanolamine (Etn) for the CDP-Etn pathway, redistribute intracellular Etn and balance the CDP-Cho and CDP-Etn arms of the Kennedy pathway. Involved in membrane synthesis and myelin production.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K90(3.72) | LDD0277 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K75(2.45) | LDD3494 | [2] | |
|
DBIA Probe Info |
 |
C159(2.94) | LDD3427 | [3] | |
|
ART-yne Probe Info |
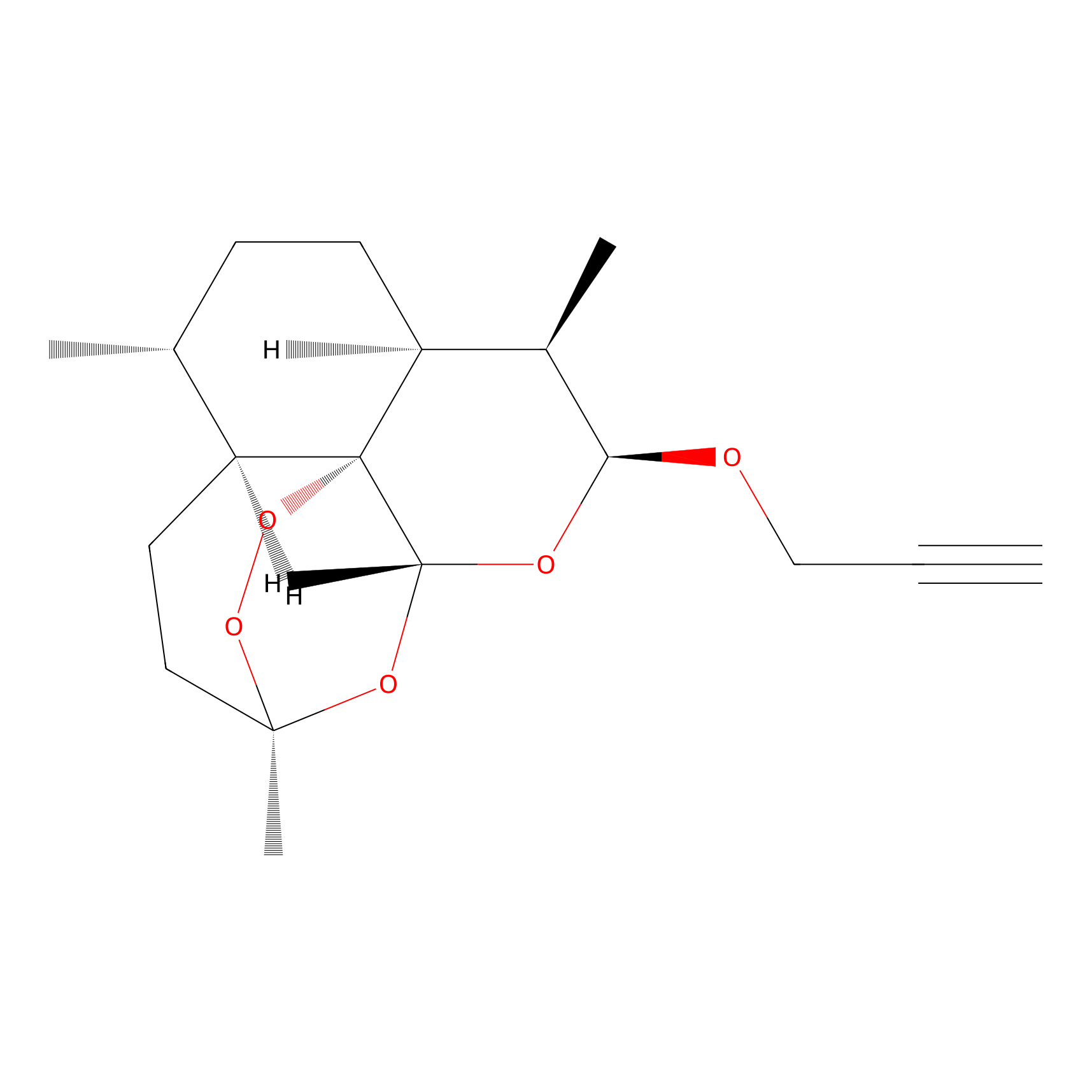 |
1.78 | LDD0234 | [4] | |
|
IA-alkyne Probe Info |
 |
C98(0.00); C159(0.00); C69(0.00) | LDD0162 | [5] | |
|
IPM Probe Info |
 |
C4(0.00); C3(0.00) | LDD2156 | [6] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C228 Probe Info |
 |
15.89 | LDD1901 | [7] | |
|
C376 Probe Info |
 |
6.73 | LDD2036 | [7] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0259 | AC14 | HEK-293T | C117(0.93) | LDD1512 | [8] |
| LDCM0282 | AC22 | HEK-293T | C117(0.92) | LDD1521 | [8] |
| LDCM0291 | AC30 | HEK-293T | C117(0.93) | LDD1530 | [8] |
| LDCM0299 | AC38 | HEK-293T | C117(0.98) | LDD1538 | [8] |
| LDCM0308 | AC46 | HEK-293T | C117(0.94) | LDD1547 | [8] |
| LDCM0317 | AC54 | HEK-293T | C117(0.93) | LDD1556 | [8] |
| LDCM0323 | AC6 | HEK-293T | C117(0.94) | LDD1562 | [8] |
| LDCM0326 | AC62 | HEK-293T | C117(0.89) | LDD1565 | [8] |
| LDCM0091 | Astemisinin | MCF-7 | 1.78 | LDD0234 | [4] |
| LDCM0368 | CL10 | HEK-293T | C117(0.74) | LDD1572 | [8] |
| LDCM0410 | CL22 | HEK-293T | C117(0.98) | LDD1614 | [8] |
| LDCM0423 | CL34 | HEK-293T | C117(1.02) | LDD1627 | [8] |
| LDCM0436 | CL46 | HEK-293T | C117(0.99) | LDD1640 | [8] |
| LDCM0449 | CL58 | HEK-293T | C117(0.90) | LDD1652 | [8] |
| LDCM0463 | CL70 | HEK-293T | C117(0.96) | LDD1666 | [8] |
| LDCM0476 | CL82 | HEK-293T | C117(0.94) | LDD1679 | [8] |
| LDCM0489 | CL94 | HEK-293T | C117(1.11) | LDD1692 | [8] |
| LDCM0572 | Fragment10 | Ramos | 8.30 | LDD2189 | [9] |
| LDCM0573 | Fragment11 | Ramos | 6.44 | LDD2190 | [9] |
| LDCM0574 | Fragment12 | Ramos | 4.55 | LDD2191 | [9] |
| LDCM0576 | Fragment14 | Ramos | 0.75 | LDD2193 | [9] |
| LDCM0580 | Fragment21 | Ramos | 2.05 | LDD2195 | [9] |
| LDCM0582 | Fragment23 | Ramos | 10.01 | LDD2196 | [9] |
| LDCM0578 | Fragment27 | Ramos | 2.58 | LDD2197 | [9] |
| LDCM0588 | Fragment30 | Ramos | 2.84 | LDD2199 | [9] |
| LDCM0468 | Fragment33 | Ramos | 4.04 | LDD2202 | [9] |
| LDCM0022 | KB02 | Ramos | 0.94 | LDD2182 | [9] |
| LDCM0023 | KB03 | Hs 936.T | C159(4.67) | LDD2780 | [3] |
| LDCM0024 | KB05 | SK-MEL-28 | C159(2.94) | LDD3427 | [3] |
The Interaction Atlas With This Target
References
