Details of the Target
General Information of Target
| Target ID | LDTP09587 | |||||
|---|---|---|---|---|---|---|
| Target Name | Protein BRICK1 (BRK1) | |||||
| Gene Name | BRK1 | |||||
| Gene ID | 55845 | |||||
| Synonyms |
C3orf10; Protein BRICK1; BRK1 |
|||||
| 3D Structure | ||||||
| Sequence |
MAGQEDPVQREIHQDWANREYIEIITSSIKKIADFLNSFDMSCRSRLATLNEKLTALERR
IEYIEARVTKGETLT |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
BRK1 family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton
|
|||||
| Function |
Involved in regulation of actin and microtubule organization. Part of a WAVE complex that activates the Arp2/3 complex. As component of the WAVE1 complex, required for BDNF-NTRK2 endocytic trafficking and signaling from early endosomes.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C43(2.62) | LDD3333 | [1] | |
|
Alkyne-RA190 Probe Info |
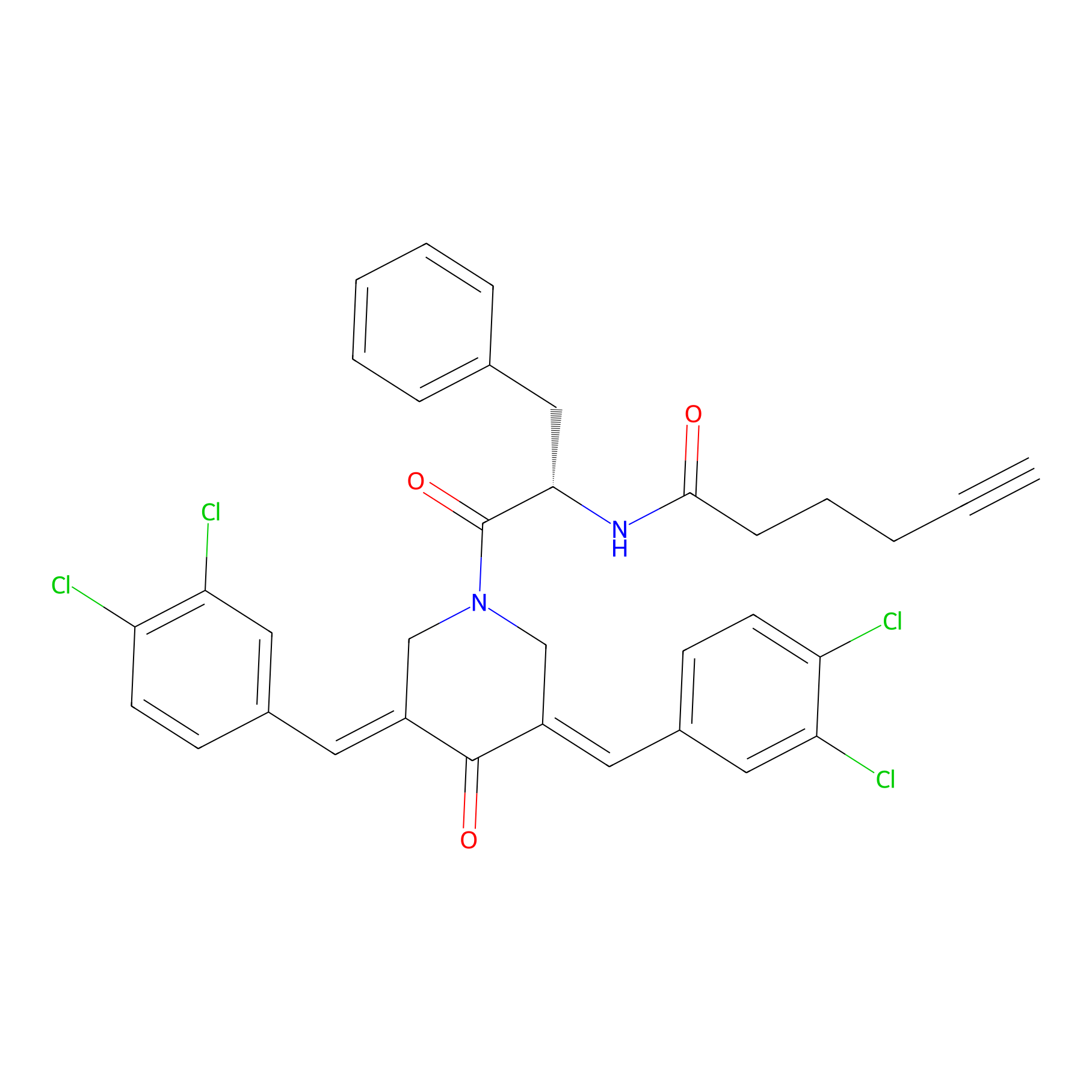 |
3.57 | LDD0299 | [2] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0224 | [3] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0175 | [4] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [5] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [6] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [5] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [7] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [8] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [7] | |
|
AOyne Probe Info |
 |
14.80 | LDD0443 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C43(0.86) | LDD1507 | [10] |
| LDCM0259 | AC14 | HEK-293T | C43(0.97) | LDD1512 | [10] |
| LDCM0276 | AC17 | HEK-293T | C43(1.00) | LDD1515 | [10] |
| LDCM0281 | AC21 | HEK-293T | C43(0.90) | LDD1520 | [10] |
| LDCM0282 | AC22 | HEK-293T | C43(0.98) | LDD1521 | [10] |
| LDCM0285 | AC25 | HEK-293T | C43(0.91) | LDD1524 | [10] |
| LDCM0289 | AC29 | HEK-293T | C43(0.97) | LDD1528 | [10] |
| LDCM0291 | AC30 | HEK-293T | C43(0.97) | LDD1530 | [10] |
| LDCM0294 | AC33 | HEK-293T | C43(1.03) | LDD1533 | [10] |
| LDCM0298 | AC37 | HEK-293T | C43(1.01) | LDD1537 | [10] |
| LDCM0299 | AC38 | HEK-293T | C43(1.03) | LDD1538 | [10] |
| LDCM0303 | AC41 | HEK-293T | C43(1.12) | LDD1542 | [10] |
| LDCM0307 | AC45 | HEK-293T | C43(1.15) | LDD1546 | [10] |
| LDCM0308 | AC46 | HEK-293T | C43(1.03) | LDD1547 | [10] |
| LDCM0311 | AC49 | HEK-293T | C43(1.11) | LDD1550 | [10] |
| LDCM0312 | AC5 | HEK-293T | C43(1.05) | LDD1551 | [10] |
| LDCM0316 | AC53 | HEK-293T | C43(1.00) | LDD1555 | [10] |
| LDCM0317 | AC54 | HEK-293T | C43(1.06) | LDD1556 | [10] |
| LDCM0320 | AC57 | HEK-293T | C43(1.05) | LDD1559 | [10] |
| LDCM0323 | AC6 | HEK-293T | C43(1.00) | LDD1562 | [10] |
| LDCM0325 | AC61 | HEK-293T | C43(0.99) | LDD1564 | [10] |
| LDCM0326 | AC62 | HEK-293T | C43(1.06) | LDD1565 | [10] |
| LDCM0248 | AKOS034007472 | HEK-293T | C43(0.97) | LDD1511 | [10] |
| LDCM0356 | AKOS034007680 | HEK-293T | C43(1.10) | LDD1570 | [10] |
| LDCM0632 | CL-Sc | Hep-G2 | C43(1.08) | LDD2227 | [6] |
| LDCM0368 | CL10 | HEK-293T | C43(1.09) | LDD1572 | [10] |
| LDCM0369 | CL100 | HEK-293T | C43(0.99) | LDD1573 | [10] |
| LDCM0372 | CL103 | HEK-293T | C43(1.10) | LDD1576 | [10] |
| LDCM0373 | CL104 | HEK-293T | C43(1.01) | LDD1577 | [10] |
| LDCM0376 | CL107 | HEK-293T | C43(1.04) | LDD1580 | [10] |
| LDCM0377 | CL108 | HEK-293T | C43(0.93) | LDD1581 | [10] |
| LDCM0381 | CL111 | HEK-293T | C43(1.20) | LDD1585 | [10] |
| LDCM0382 | CL112 | HEK-293T | C43(0.88) | LDD1586 | [10] |
| LDCM0385 | CL115 | HEK-293T | C43(1.13) | LDD1589 | [10] |
| LDCM0386 | CL116 | HEK-293T | C43(0.94) | LDD1590 | [10] |
| LDCM0389 | CL119 | HEK-293T | C43(1.18) | LDD1593 | [10] |
| LDCM0391 | CL120 | HEK-293T | C43(1.00) | LDD1595 | [10] |
| LDCM0394 | CL123 | HEK-293T | C43(1.35) | LDD1598 | [10] |
| LDCM0395 | CL124 | HEK-293T | C43(1.13) | LDD1599 | [10] |
| LDCM0398 | CL127 | HEK-293T | C43(1.21) | LDD1602 | [10] |
| LDCM0399 | CL128 | HEK-293T | C43(1.08) | LDD1603 | [10] |
| LDCM0402 | CL15 | HEK-293T | C43(1.20) | LDD1606 | [10] |
| LDCM0403 | CL16 | HEK-293T | C43(0.95) | LDD1607 | [10] |
| LDCM0404 | CL17 | HEK-293T | C43(1.09) | LDD1608 | [10] |
| LDCM0409 | CL21 | HEK-293T | C43(0.87) | LDD1613 | [10] |
| LDCM0410 | CL22 | HEK-293T | C43(1.04) | LDD1614 | [10] |
| LDCM0415 | CL27 | HEK-293T | C43(1.08) | LDD1619 | [10] |
| LDCM0416 | CL28 | HEK-293T | C43(0.98) | LDD1620 | [10] |
| LDCM0417 | CL29 | HEK-293T | C43(0.98) | LDD1621 | [10] |
| LDCM0418 | CL3 | HEK-293T | C43(1.15) | LDD1622 | [10] |
| LDCM0422 | CL33 | HEK-293T | C43(0.96) | LDD1626 | [10] |
| LDCM0423 | CL34 | HEK-293T | C43(0.96) | LDD1627 | [10] |
| LDCM0428 | CL39 | HEK-293T | C43(1.04) | LDD1632 | [10] |
| LDCM0429 | CL4 | HEK-293T | C43(1.01) | LDD1633 | [10] |
| LDCM0430 | CL40 | HEK-293T | C43(0.96) | LDD1634 | [10] |
| LDCM0431 | CL41 | HEK-293T | C43(0.89) | LDD1635 | [10] |
| LDCM0435 | CL45 | HEK-293T | C43(0.87) | LDD1639 | [10] |
| LDCM0436 | CL46 | HEK-293T | C43(1.06) | LDD1640 | [10] |
| LDCM0440 | CL5 | HEK-293T | C43(0.91) | LDD1644 | [10] |
| LDCM0443 | CL52 | HEK-293T | C43(0.95) | LDD1646 | [10] |
| LDCM0444 | CL53 | HEK-293T | C43(1.17) | LDD1647 | [10] |
| LDCM0448 | CL57 | HEK-293T | C43(0.84) | LDD1651 | [10] |
| LDCM0449 | CL58 | HEK-293T | C43(0.91) | LDD1652 | [10] |
| LDCM0455 | CL63 | HEK-293T | C43(1.03) | LDD1658 | [10] |
| LDCM0456 | CL64 | HEK-293T | C43(1.18) | LDD1659 | [10] |
| LDCM0457 | CL65 | HEK-293T | C43(1.05) | LDD1660 | [10] |
| LDCM0461 | CL69 | HEK-293T | C43(0.78) | LDD1664 | [10] |
| LDCM0463 | CL70 | HEK-293T | C43(0.87) | LDD1666 | [10] |
| LDCM0469 | CL76 | HEK-293T | C43(0.92) | LDD1672 | [10] |
| LDCM0470 | CL77 | HEK-293T | C43(0.97) | LDD1673 | [10] |
| LDCM0475 | CL81 | HEK-293T | C43(0.80) | LDD1678 | [10] |
| LDCM0476 | CL82 | HEK-293T | C43(1.10) | LDD1679 | [10] |
| LDCM0481 | CL87 | HEK-293T | C43(1.09) | LDD1684 | [10] |
| LDCM0482 | CL88 | HEK-293T | C43(0.98) | LDD1685 | [10] |
| LDCM0483 | CL89 | HEK-293T | C43(1.04) | LDD1686 | [10] |
| LDCM0484 | CL9 | HEK-293T | C43(0.78) | LDD1687 | [10] |
| LDCM0488 | CL93 | HEK-293T | C43(0.93) | LDD1691 | [10] |
| LDCM0489 | CL94 | HEK-293T | C43(1.25) | LDD1692 | [10] |
| LDCM0494 | CL99 | HEK-293T | C43(0.82) | LDD1697 | [10] |
| LDCM0495 | E2913 | HEK-293T | C43(0.68) | LDD1698 | [10] |
| LDCM0625 | F8 | Ramos | C43(0.47) | LDD2187 | [11] |
| LDCM0572 | Fragment10 | Ramos | C43(1.05) | LDD2189 | [11] |
| LDCM0573 | Fragment11 | Ramos | C43(0.17) | LDD2190 | [11] |
| LDCM0574 | Fragment12 | Ramos | C43(1.07) | LDD2191 | [11] |
| LDCM0575 | Fragment13 | Ramos | C43(0.78) | LDD2192 | [11] |
| LDCM0576 | Fragment14 | Ramos | C43(1.75) | LDD2193 | [11] |
| LDCM0579 | Fragment20 | Ramos | C43(0.82) | LDD2194 | [11] |
| LDCM0580 | Fragment21 | Ramos | C43(0.77) | LDD2195 | [11] |
| LDCM0582 | Fragment23 | Ramos | C43(0.87) | LDD2196 | [11] |
| LDCM0578 | Fragment27 | Ramos | C43(0.70) | LDD2197 | [11] |
| LDCM0586 | Fragment28 | Ramos | C43(0.56) | LDD2198 | [11] |
| LDCM0588 | Fragment30 | Ramos | C43(1.45) | LDD2199 | [11] |
| LDCM0589 | Fragment31 | Ramos | C43(0.86) | LDD2200 | [11] |
| LDCM0590 | Fragment32 | Ramos | C43(0.79) | LDD2201 | [11] |
| LDCM0468 | Fragment33 | HEK-293T | C43(1.05) | LDD1671 | [10] |
| LDCM0596 | Fragment38 | Ramos | C43(1.02) | LDD2203 | [11] |
| LDCM0566 | Fragment4 | Ramos | C43(1.53) | LDD2184 | [11] |
| LDCM0610 | Fragment52 | Ramos | C43(0.95) | LDD2204 | [11] |
| LDCM0614 | Fragment56 | Ramos | C43(0.85) | LDD2205 | [11] |
| LDCM0569 | Fragment7 | Ramos | C43(0.73) | LDD2186 | [11] |
| LDCM0571 | Fragment9 | Ramos | C43(0.72) | LDD2188 | [11] |
| LDCM0022 | KB02 | HEK-293T | C43(0.97) | LDD1492 | [10] |
| LDCM0023 | KB03 | HEK-293T | C43(1.13) | LDD1497 | [10] |
| LDCM0024 | KB05 | MOLM-13 | C43(2.62) | LDD3333 | [1] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [3] |
| LDCM0131 | RA190 | MM1.R | 3.57 | LDD0299 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Transcription factor
GPCR
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| P2Y purinoceptor 2 (P2RY2) | G-protein coupled receptor 1 family | P41231 | |||
Other
References
