Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | SNV: p.C106Y | . | |||
| AN3CA | SNV: p.G1775E | DBIA Probe Info | |||
| CAL51 | SNV: p.G766D | DBIA Probe Info | |||
| CASKI | SNV: p.M1513I | DBIA Probe Info | |||
| DETROIT562 | SNV: p.Q1018R | . | |||
| FTC133 | SNV: p.A95T | DBIA Probe Info | |||
| G361 | SNV: p.R982C | DBIA Probe Info | |||
| HCT116 | Insertion: p.E653Ter | . | |||
| HCT15 | SNV: p.N813S | . | |||
| HEC1B | SNV: p.S998N | . | |||
| HT | SNV: p.K759N | . | |||
| HT115 | SNV: p.T1748M | . | |||
| ICC3 | SNV: p.R187Q | DBIA Probe Info | |||
| IGR37 | SNV: p.P750S | . | |||
| IGR39 | SNV: p.P750S | DBIA Probe Info | |||
| MELJUSO | SNV: p.L1350Q | DBIA Probe Info | |||
| NB1 | SNV: p.A50D; p.E1790A | . | |||
| NCIH23 | SNV: p.V278L | DBIA Probe Info | |||
| OV90 | SNV: p.M1513V | DBIA Probe Info | |||
| OVCAR8 | SNV: p.G902V | DBIA Probe Info | |||
| OVK18 | SNV: p.N1324T | DBIA Probe Info | |||
| PC14 | SNV: p.Y1353S | . | |||
| RKO | SNV: p.V199D | DBIA Probe Info | |||
| SNGM | SNV: p.G847V; p.A1127T; p.F1463V | DBIA Probe Info | |||
| U87MG | SNV: p.A1762V | DBIA Probe Info | |||
| UMUC3 | SNV: p.S13Y | DBIA Probe Info | |||
| ZR751 | SNV: p.E1414K | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
IPM Probe Info |
 |
N.A. | LDD0241 | [1] | |
|
DBIA Probe Info |
 |
C1305(1.16); C1307(1.16); C828(0.99); C832(0.99) | LDD0078 | [2] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [3] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [1] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DFG-out-4 Probe Info |
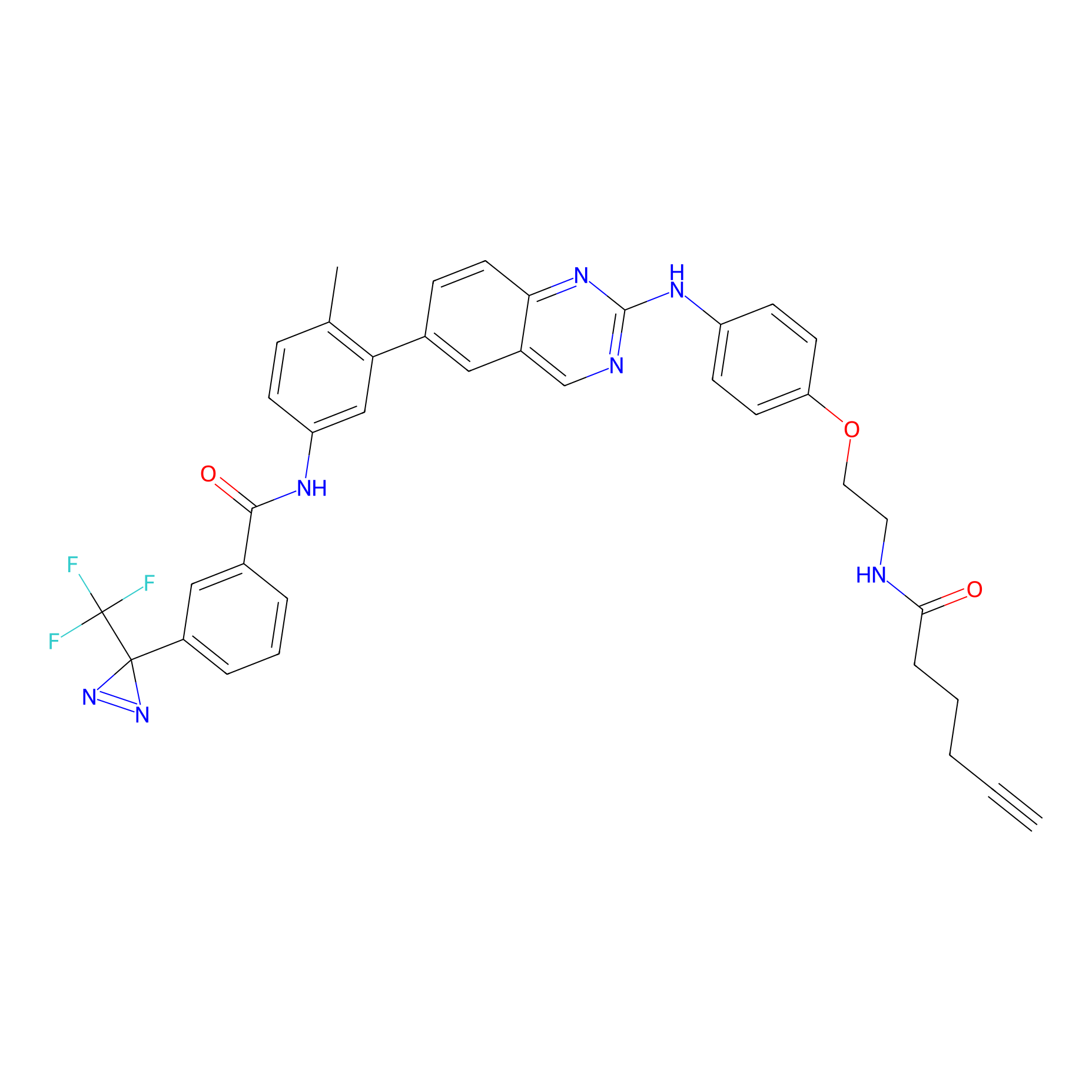 |
9.20 | LDD0075 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0259 | AC14 | HEK-293T | C297(1.13) | LDD1512 | [5] |
| LDCM0282 | AC22 | HEK-293T | C297(0.70) | LDD1521 | [5] |
| LDCM0284 | AC24 | HEK-293T | C435(0.96) | LDD1523 | [5] |
| LDCM0291 | AC30 | HEK-293T | C297(1.05) | LDD1530 | [5] |
| LDCM0293 | AC32 | HEK-293T | C435(0.98) | LDD1532 | [5] |
| LDCM0299 | AC38 | HEK-293T | C297(0.84) | LDD1538 | [5] |
| LDCM0302 | AC40 | HEK-293T | C435(1.10) | LDD1541 | [5] |
| LDCM0308 | AC46 | HEK-293T | C297(1.28) | LDD1547 | [5] |
| LDCM0310 | AC48 | HEK-293T | C435(1.13) | LDD1549 | [5] |
| LDCM0317 | AC54 | HEK-293T | C297(1.06) | LDD1556 | [5] |
| LDCM0319 | AC56 | HEK-293T | C435(1.02) | LDD1558 | [5] |
| LDCM0323 | AC6 | HEK-293T | C297(1.14) | LDD1562 | [5] |
| LDCM0326 | AC62 | HEK-293T | C297(0.93) | LDD1565 | [5] |
| LDCM0328 | AC64 | HEK-293T | C435(1.04) | LDD1567 | [5] |
| LDCM0345 | AC8 | HEK-293T | C435(1.12) | LDD1569 | [5] |
| LDCM0275 | AKOS034007705 | HEK-293T | C435(1.05) | LDD1514 | [5] |
| LDCM0020 | ARS-1620 | HCC44 | C1305(1.16); C1307(1.16); C828(0.99); C832(0.99) | LDD0078 | [2] |
| LDCM0367 | CL1 | HEK-293T | C297(0.99) | LDD1571 | [5] |
| LDCM0368 | CL10 | HEK-293T | C297(0.51) | LDD1572 | [5] |
| LDCM0370 | CL101 | HEK-293T | C297(0.96) | LDD1574 | [5] |
| LDCM0374 | CL105 | HEK-293T | C297(1.10) | LDD1578 | [5] |
| LDCM0378 | CL109 | HEK-293T | C297(1.23) | LDD1582 | [5] |
| LDCM0383 | CL113 | HEK-293T | C297(1.21) | LDD1587 | [5] |
| LDCM0387 | CL117 | HEK-293T | C297(0.94) | LDD1591 | [5] |
| LDCM0390 | CL12 | HEK-293T | C435(1.64) | LDD1594 | [5] |
| LDCM0392 | CL121 | HEK-293T | C297(1.40) | LDD1596 | [5] |
| LDCM0396 | CL125 | HEK-293T | C297(0.99) | LDD1600 | [5] |
| LDCM0400 | CL13 | HEK-293T | C297(1.22) | LDD1604 | [5] |
| LDCM0410 | CL22 | HEK-293T | C297(1.05) | LDD1614 | [5] |
| LDCM0412 | CL24 | HEK-293T | C435(1.53) | LDD1616 | [5] |
| LDCM0413 | CL25 | HEK-293T | C297(0.96) | LDD1617 | [5] |
| LDCM0423 | CL34 | HEK-293T | C297(1.65) | LDD1627 | [5] |
| LDCM0425 | CL36 | HEK-293T | C435(1.19) | LDD1629 | [5] |
| LDCM0426 | CL37 | HEK-293T | C297(0.92) | LDD1630 | [5] |
| LDCM0436 | CL46 | HEK-293T | C297(0.90) | LDD1640 | [5] |
| LDCM0438 | CL48 | HEK-293T | C435(1.44) | LDD1642 | [5] |
| LDCM0439 | CL49 | HEK-293T | C297(0.92) | LDD1643 | [5] |
| LDCM0449 | CL58 | HEK-293T | C297(1.04) | LDD1652 | [5] |
| LDCM0452 | CL60 | HEK-293T | C435(1.24) | LDD1655 | [5] |
| LDCM0453 | CL61 | HEK-293T | C297(1.26) | LDD1656 | [5] |
| LDCM0463 | CL70 | HEK-293T | C297(0.97) | LDD1666 | [5] |
| LDCM0465 | CL72 | HEK-293T | C435(1.20) | LDD1668 | [5] |
| LDCM0466 | CL73 | HEK-293T | C297(0.96) | LDD1669 | [5] |
| LDCM0476 | CL82 | HEK-293T | C297(1.18) | LDD1679 | [5] |
| LDCM0478 | CL84 | HEK-293T | C435(1.66) | LDD1681 | [5] |
| LDCM0479 | CL85 | HEK-293T | C297(0.85) | LDD1682 | [5] |
| LDCM0489 | CL94 | HEK-293T | C297(0.88) | LDD1692 | [5] |
| LDCM0491 | CL96 | HEK-293T | C435(1.53) | LDD1694 | [5] |
| LDCM0492 | CL97 | HEK-293T | C297(0.83) | LDD1695 | [5] |
| LDCM0017 | DFG-out-2 | A431 | 9.20 | LDD0075 | [4] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C473(1.41); C297(0.90) | LDD1702 | [6] |
| LDCM0022 | KB02 | HEK-293T | C83(0.94) | LDD1492 | [5] |
| LDCM0023 | KB03 | HEK-293T | C83(0.96) | LDD1497 | [5] |
| LDCM0024 | KB05 | COLO792 | C462(2.06); C83(1.79); C155(1.58) | LDD3310 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Golgi-associated PDZ and coiled-coil motif-containing protein (GOPC) | . | Q9HD26 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amyloid protein-binding protein 2 (APPBP2) | . | Q92624 | |||
| SH2/SH3 adapter protein NCK1 (NCK1) | . | P16333 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Fostamatinib | Small molecular drug | DB12010 | |||
| Ritodrine | Small molecular drug | DB00867 | |||
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Me-3407 | Small molecular drug | D0L8MO | |||
Investigative
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Rki-1447 | Small molecular drug | D03CPB | |||
References
