Details of the Target
General Information of Target
| Target ID | LDTP04717 | |||||
|---|---|---|---|---|---|---|
| Target Name | Small ubiquitin-related modifier 3 (SUMO3) | |||||
| Gene Name | SUMO3 | |||||
| Gene ID | 6612 | |||||
| Synonyms |
SMT3A; SMT3H1; Small ubiquitin-related modifier 3; SUMO-3; SMT3 homolog 1; SUMO-2; Ubiquitin-like protein SMT3A; Smt3A |
|||||
| 3D Structure | ||||||
| Sequence |
MSEEKPKEGVKTENDHINLKVAGQDGSVVQFKIKRHTPLSKLMKAYCERQGLSMRQIRFR
FDGQPINETDTPAQLEMEDEDTIDVFQQQTGGVPESSLAGHSF |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Ubiquitin family, SUMO subfamily
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Ubiquitin-like protein which can be covalently attached to target lysines either as a monomer or as a lysine-linked polymer. Does not seem to be involved in protein degradation and may function as an antagonist of ubiquitin in the degradation process. Plays a role in a number of cellular processes such as nuclear transport, DNA replication and repair, mitosis and signal transduction. Covalent attachment to its substrates requires prior activation by the E1 complex SAE1-SAE2 and linkage to the E2 enzyme UBE2I, and can be promoted by an E3 ligase such as PIAS1-4, RANBP2 or CBX4. Plays a role in the regulation of sumoylation status of SETX.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
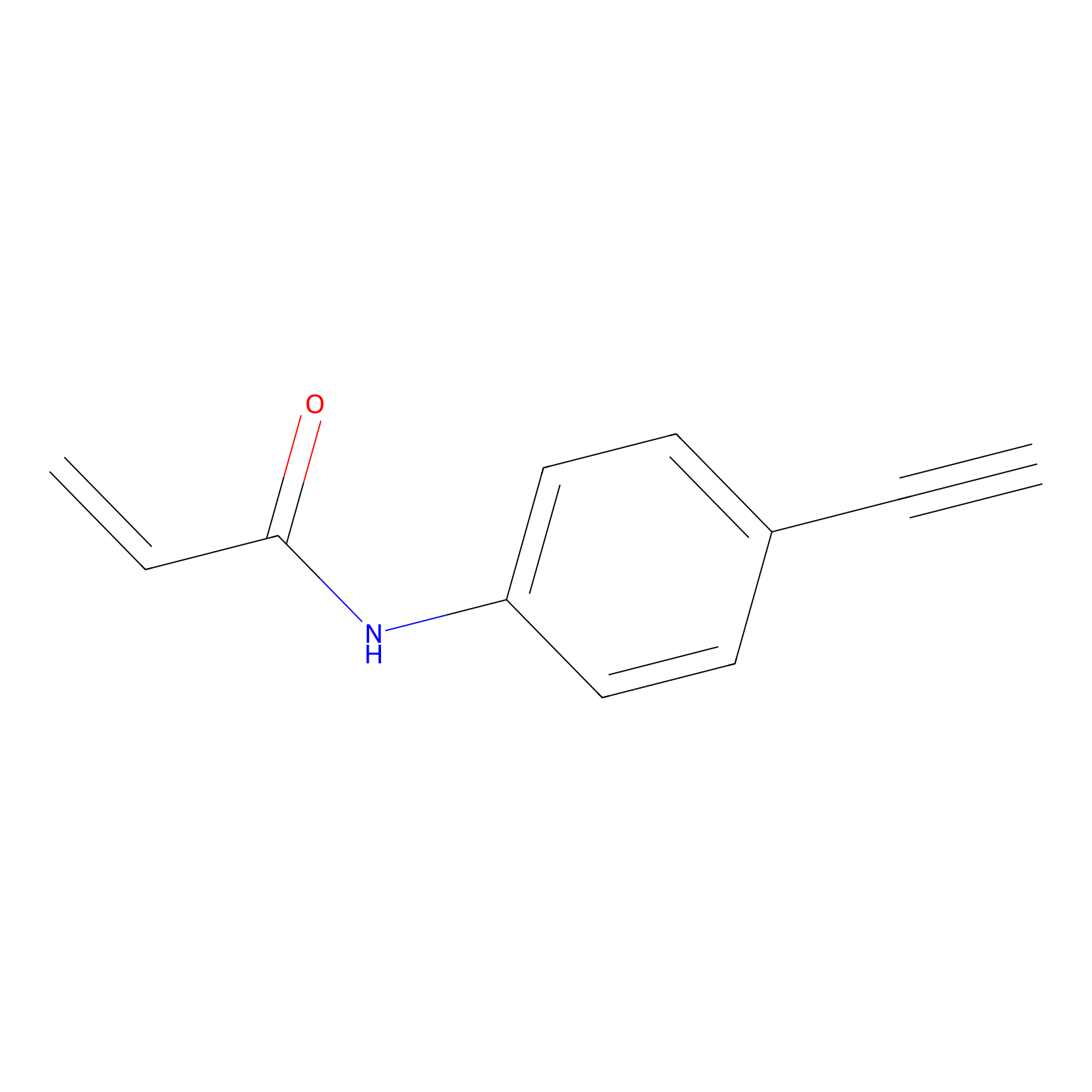 |
2.10 | LDD0449 | [1] | |
|
P3 Probe Info |
 |
1.82 | LDD0450 | [1] | |
|
ONAyne Probe Info |
 |
K32(1.02) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K32(4.46); K41(5.88); K44(5.26) | LDD0277 | [2] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0221 | [3] | |
|
AMP probe Probe Info |
 |
K32(0.00); K34(0.00) | LDD0200 | [4] | |
|
ATP probe Probe Info |
 |
K32(0.00); K34(0.00); K41(0.00); K11(0.00) | LDD0199 | [4] | |
|
ATP probe Probe Info |
 |
K32(0.00); K41(0.00) | LDD0035 | [5] | |
|
HHS-465 Probe Info |
 |
K32(0.00); K44(0.00); Y46(0.00); K41(0.00) | LDD2240 | [6] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| SUMO-specific isopeptidase USPL1 (USPL1) | Peptidase C19 family | Q5W0Q7 | |||
| Sentrin-specific protease 2 (SENP2) | Peptidase C48 family | Q9HC62 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Huntingtin (HTT) | Huntingtin family | P42858 | |||
Transcription factor
Other
References
