Details of the Target
General Information of Target
| Target ID | LDTP02452 | |||||
|---|---|---|---|---|---|---|
| Target Name | Tubulin alpha-3C chain (TUBA3C) | |||||
| Gene Name | TUBA3C | |||||
| Gene ID | 7278 | |||||
| Synonyms |
TUBA2; Tubulin alpha-3C chain; EC 3.6.5.-; Alpha-tubulin 2; Alpha-tubulin 3C; Tubulin alpha-2 chain) [Cleaved into: Detyrosinated tubulin alpha-3C chain] |
|||||
| 3D Structure | ||||||
| Sequence |
MRECISIHVGQAGVQIGNACWELYCLEHGIQPDGQMPSDKTIGGGDDSFNTFFSETGAGK
HVPRAVFVDLEPTVVDEVRTGTYRQLFHPEQLITGKEDAANNYARGHYTIGKEIVDLVLD RIRKLADLCTGLQGFLIFHSFGGGTGSGFASLLMERLSVDYGKKSKLEFAIYPAPQVSTA VVEPYNSILTTHTTLEHSDCAFMVDNEAIYDICRRNLDIERPTYTNLNRLIGQIVSSITA SLRFDGALNVDLTEFQTNLVPYPRIHFPLATYAPVISAEKAYHEQLSVAEITNACFEPAN QMVKCDPRHGKYMACCMLYRGDVVPKDVNAAIATIKTKRTIQFVDWCPTGFKVGINYQPP TVVPGGDLAKVQRAVCMLSNTTAIAEAWARLDHKFDLMYAKRAFVHWYVGEGMEEGEFSE AREDLAALEKDYEEVGVDSVEAEAEEGEEY |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Tubulin family
|
|||||
| Subcellular location |
Cytoplasm, cytoskeleton
|
|||||
| Function |
Tubulin is the major constituent of microtubules, a cylinder consisting of laterally associated linear protofilaments composed of alpha- and beta-tubulin heterodimers. Microtubules grow by the addition of GTP-tubulin dimers to the microtubule end, where a stabilizing cap forms. Below the cap, tubulin dimers are in GDP-bound state, owing to GTPase activity of alpha-tubulin.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
JZ128-DTB Probe Info |
 |
C295(0.00); C376(0.00); C305(0.00) | LDD0462 | [1] | |
|
THZ1-DTB Probe Info |
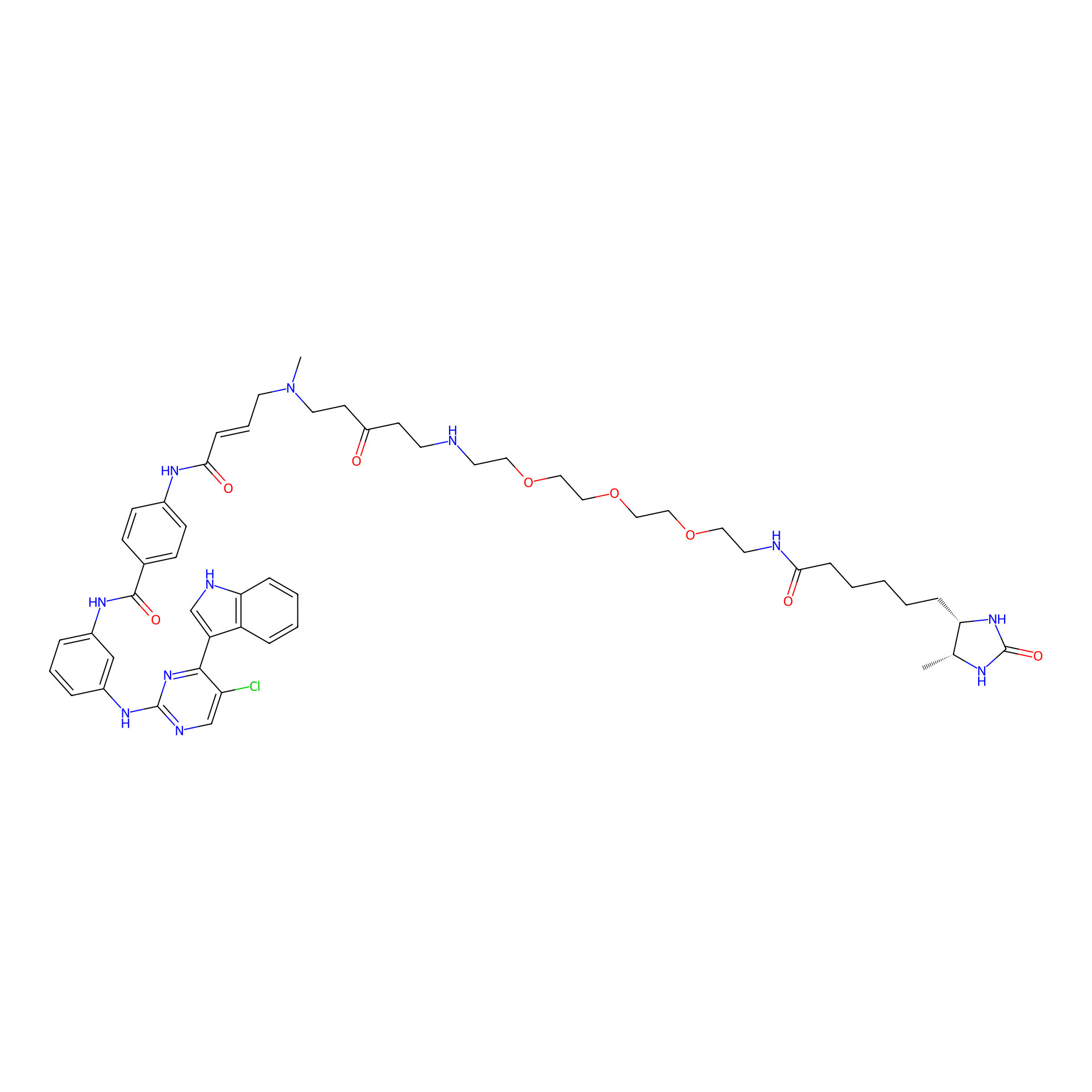 |
C347(1.14); C376(1.06); C295(1.05); C25(1.06) | LDD0460 | [1] | |
|
HHS-475 Probe Info |
 |
Y103(0.64); Y399(0.65); Y262(0.67); Y272(0.77) | LDD0264 | [2] | |
|
5E-2FA Probe Info |
 |
H283(0.00); H266(0.00); H88(0.00); H61(0.00) | LDD2235 | [3] | |
|
m-APA Probe Info |
 |
H406(0.00); H283(0.00); H266(0.00); H88(0.00) | LDD2231 | [3] | |
|
Ox-W18 Probe Info |
 |
W346(0.00); W388(0.00) | LDD2175 | [4] | |
|
NAIA_5 Probe Info |
 |
C376(0.00); C305(0.00); C347(0.00); C4(0.00) | LDD2223 | [5] | |
|
HHS-465 Probe Info |
 |
K280(0.00); K326(0.00); K336(0.00); K370(0.00) | LDD2240 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0116 | HHS-0101 | DM93 | Y103(0.64); Y399(0.65); Y262(0.67); Y272(0.77) | LDD0264 | [2] |
| LDCM0117 | HHS-0201 | DM93 | Y399(0.53); Y161(0.53); Y103(0.60); Y262(0.60) | LDD0265 | [2] |
| LDCM0118 | HHS-0301 | DM93 | Y262(0.52); Y399(0.61); Y272(0.64); Y103(0.67) | LDD0266 | [2] |
| LDCM0119 | HHS-0401 | DM93 | Y272(0.39); Y262(0.44); Y399(0.49); Y103(0.54) | LDD0267 | [2] |
| LDCM0120 | HHS-0701 | DM93 | Y103(0.40); Y272(0.47); Y262(0.52); Y399(0.55) | LDD0268 | [2] |
| LDCM0179 | JZ128 | PC-3 | C295(0.00); C376(0.00); C305(0.00) | LDD0462 | [1] |
| LDCM0021 | THZ1 | HeLa S3 | C347(1.14); C376(1.06); C295(1.05); C25(1.06) | LDD0460 | [1] |
References
