Details of the Target
General Information of Target
| Target ID | LDTP01611 | |||||
|---|---|---|---|---|---|---|
| Target Name | Formin-like protein 1 (FMNL1) | |||||
| Gene Name | FMNL1 | |||||
| Gene ID | 752 | |||||
| Synonyms |
C17orf1; C17orf1B; FMNL; FRL1; Formin-like protein 1; CLL-associated antigen KW-13; Leukocyte formin |
|||||
| 3D Structure | ||||||
| Sequence |
MGNAAGSAEQPAGPAAPPPKQPAPPKQPMPAAGELEERFNRALNCMNLPPDKVQLLSQYD
NEKKWELICDQERFQVKNPPAAYIQKLKSYVDTGGVSRKVAADWMSNLGFKRRVQESTQV LRELETSLRTNHIGWVQEFLNEENRGLDVLLEYLAFAQCSVTYDMESTDNGASNSEKNKP LEQSVEDLSKGPPSSVPKSRHLTIKLTPAHSRKALRNSRIVSQKDDVHVCIMCLRAIMNY QSGFSLVMNHPACVNEIALSLNNKNPRTKALVLELLAAVCLVRGGHDIILAAFDNFKEVC GEQHRFEKLMEYFRNEDSNIDFMVACMQFINIVVHSVENMNFRVFLQYEFTHLGLDLYLE RLRLTESDKLQVQIQAYLDNIFDVGALLEDTETKNAVLEHMEELQEQVALLTERLRDAEN ESMAKIAELEKQLSQARKELETLRERFSESTAMGPSRRPPEPEKAPPAAPTRPSALELKV EELEEKGLIRILRGPGDAVSIEILPVAVATPSGGDAPTPGVPTGSPSPDLAPAAEPAPGA APPPPPPLPGLPSPQEAPPSAPPQAPPLPGSPEPPPAPPLPGDLPPPPPPPPPPPGTDGP VPPPPPPPPPPPGGPPDALGRRDSELGPGVKAKKPIQTKFRMPLLNWVALKPSQITGTVF TELNDEKVLQELDMSDFEEQFKTKSQGPSLDLSALKSKAAQKAPSKATLIEANRAKNLAI TLRKGNLGAERICQAIEAYDLQALGLDFLELLMRFLPTEYERSLITRFEREQRPMEELSE EDRFMLCFSRIPRLPERMTTLTFLGNFPDTAQLLMPQLNAIIAASMSIKSSDKLRQILEI VLAFGNYMNSSKRGAAYGFRLQSLDALLEMKSTDRKQTLLHYLVKVIAEKYPQLTGFHSD LHFLDKAGSVSLDSVLADVRSLQRGLELTQREFVRQDDCMVLKEFLRANSPTMDKLLADS KTAQEAFESVVEYFGENPKTTSPGLFFSLFSRFIKAYKKAEQEVEQWKKEAAAQEAGADT PGKGEPPAPKSPPKARRPQMDLISELKRRQQKEPLIYESDRDGAIEDIITVIKTVPFTAR TGKRTSRLLCEASLGEEMPL |
|||||
| Target Type |
Literature-reported
|
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Formin homology family
|
|||||
| Subcellular location |
Cytoplasm; Cytoplasm, cell cortex
|
|||||
| Function |
May play a role in the control of cell motility and survival of macrophages. Plays a role in the regulation of cell morphology and cytoskeletal organization. Required in the cortical actin filament dynamics and cell shape.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| A2058 | SNV: p.M46I | DBIA Probe Info | |||
| CAMA1 | SNV: p.S560F | DBIA Probe Info | |||
| DOTC24510 | SNV: p.E965G | . | |||
| HCT116 | Deletion: p.S527AfsTer93 | . | |||
| HT | SNV: p.M46T | DBIA Probe Info | |||
| IGROV1 | SNV: p.Q680P | . | |||
| Ishikawa (Heraklio) 02 ER | SNV: p.G243D | DBIA Probe Info | |||
| LS180 | Deletion: p.A729PfsTer24 | DBIA Probe Info | |||
| MCC13 | SNV: p.S336L | DBIA Probe Info | |||
| MOLT4 | SNV: p.S184F; p.H304Y; p.A507T; p.L756M; p.A866E; p.R920H | IA-alkyne Probe Info | |||
| NALM6 | SNV: p.F39L | DBIA Probe Info | |||
| NCIH1155 | SNV: p.R1061C | DBIA Probe Info | |||
| OVISE | SNV: p.Q564P | . | |||
| SNGM | SNV: p.A649T | DBIA Probe Info | |||
| SNU5 | SNV: p.P543Q | DBIA Probe Info | |||
| SUPT1 | SNV: p.R414W | DBIA Probe Info | |||
| TF1 | SNV: p.P545T | DBIA Probe Info | |||
| TOV21G | SNV: p.S474Ter; p.M798V | DBIA Probe Info | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y90(10.61) | LDD0260 | [1] | |
|
ONAyne Probe Info |
 |
K702(0.88) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K1047(10.00); K111(2.00); K696(10.00); K77(0.63) | LDD0277 | [2] | |
|
BTD Probe Info |
 |
C300(0.43) | LDD2091 | [3] | |
|
HPAP Probe Info |
 |
3.15 | LDD0064 | [4] | |
|
Alkyne-RA190 Probe Info |
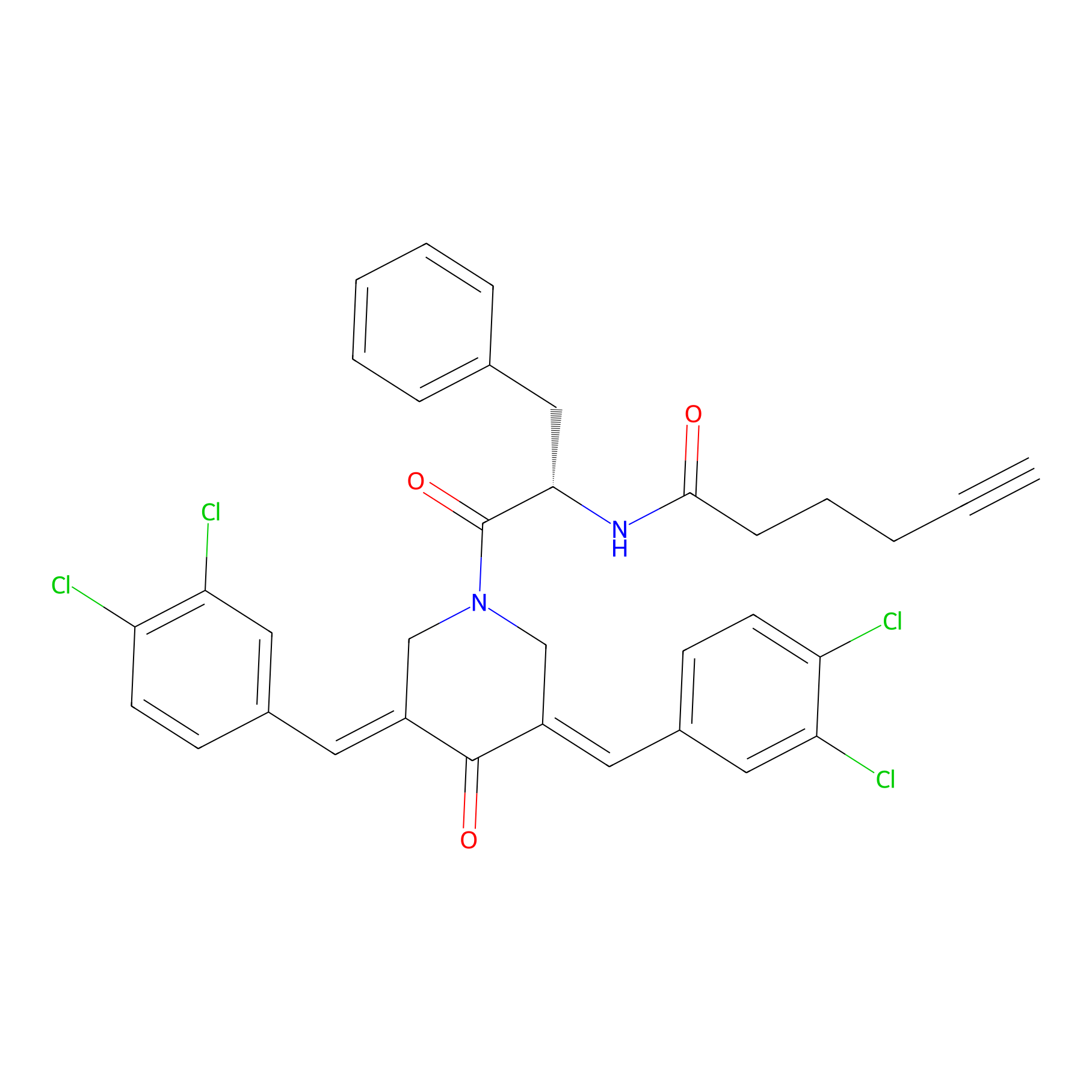 |
2.09 | LDD0300 | [5] | |
|
DBIA Probe Info |
 |
C787(8.19); C787(199.02); C300(14.46) | LDD0209 | [6] | |
|
IPM Probe Info |
 |
C69(1.95) | LDD1702 | [3] | |
|
N1 Probe Info |
 |
N.A. | LDD0245 | [7] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C253(0.00); C69(0.00); C787(0.00); C939(0.00) | LDD0038 | [8] | |
|
IA-alkyne Probe Info |
 |
C253(0.00); C787(0.00); C69(0.00); C939(0.00) | LDD0036 | [8] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
C45(0.00); C69(0.00); C253(0.00); C787(0.00) | LDD0037 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C300(1.21) | LDD2117 | [3] |
| LDCM0498 | BS-3668 | MDA-MB-231 | C300(0.43) | LDD2091 | [3] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C69(1.95) | LDD1702 | [3] |
| LDCM0022 | KB02 | T cell | C939(12.25) | LDD1707 | [10] |
| LDCM0023 | KB03 | Jurkat | C787(8.19); C787(199.02); C300(14.46) | LDD0209 | [6] |
| LDCM0024 | KB05 | IGR37 | C300(1.51) | LDD3314 | [11] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C300(1.28) | LDD2099 | [3] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C300(0.67) | LDD2104 | [3] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C300(0.90) | LDD2109 | [3] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C300(0.95) | LDD2111 | [3] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C300(0.95) | LDD2125 | [3] |
| LDCM0534 | Nucleophilic fragment 30a | MDA-MB-231 | C300(1.06) | LDD2127 | [3] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C300(0.94) | LDD2129 | [3] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C300(0.79) | LDD2133 | [3] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C300(0.58) | LDD2134 | [3] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C300(1.14) | LDD2137 | [3] |
| LDCM0552 | Nucleophilic fragment 6a | MDA-MB-231 | C300(1.11) | LDD2146 | [3] |
| LDCM0014 | Panhematin | hPBMC | 3.15 | LDD0064 | [4] |
| LDCM0131 | RA190 | MM1.R | 2.09 | LDD0300 | [5] |
The Interaction Atlas With This Target
References
