Details of the Target
General Information of Target
| Target ID | LDTP17268 | |||||
|---|---|---|---|---|---|---|
| Target Name | ATP synthase subunit epsilon-like protein, mitochondrial (ATP5F1EP2) | |||||
| Gene Name | ATP5F1EP2 | |||||
| Synonyms |
ATP5EP2; ATP synthase subunit epsilon-like protein, mitochondrial; ATP synthase F1 subunit epsilon pseudogene 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MFEDVFSDSGNTGNFDRGKKRRLTIIECGCDINMMIDLAKVADLVLMLIDASFGFEMEMF
EFLNICQAHGFPKILGVLTHLDSFKHNKQLKKTKKRLKHRFWTEVYQDKVGLTHELVQSL ISTYSTIDAKMASSRVTLLSNSKPLGSEAIDNQGVSLEFDQQQGSVCPSESEIYEAGAED RMAGAPMAAAVQPAEVTVEVGEDLHMHQVRDREMPEVVEIRRSNCTNHCDLGDTSSYHTK VSTVHIMKKRNGGGSLNNYSSSIPPTPSTSQEDPQFSVPPTANTPTPVCKRSMRWSNLFT SEKGSDPDKERKAPENHADTIGSGRAIPIKQGMLLKRSGKWLKTWKKKYVTLCSNGVLTY YSSLGDYMKNIHKKEIDLRTSTIKVPGKWPSLATSACAPISSSKSNGLSKDMDTGLGDSI CFSPGISSTTSPKLNPPPSPHANKKKHLKKKSTNNFMIVSATGQTWHFEATTYEERDAWV QAIQSQILASLQSCKSSKSKSQLTSQSEAMALQSIQNMRGNAHCVDCETQNPKWASLNLG VLMCIECSGIHRSFGTRLSRVRSLELDDWPVELRKVMSSIGNELANSIWEGSSQGQTKPS IKSTREEKEWWIRSKYEEKLFLAPLPCTELSLGQQLLRATTDEDLQTAILLLAHGSREEV NETCGEGDGCTALHLACRKGNVVLEQLLTGWTSWPEMPTGTQR |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Eukaryotic ATPase epsilon family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Mitochondrial membrane ATP synthase (F(1)F(0) ATP synthase or Complex V) produces ATP from ADP in the presence of a proton gradient across the membrane which is generated by electron transport complexes of the respiratory chain. F-type ATPases consist of two structural domains, F(1) - containing the extramembraneous catalytic core, and F(0) - containing the membrane proton channel, linked together by a central stalk and a peripheral stalk. During catalysis, ATP synthesis in the catalytic domain of F(1) is coupled via a rotary mechanism of the central stalk subunits to proton translocation. Part of the complex F(1) domain and of the central stalk which is part of the complex rotary element. Rotation of the central stalk against the surrounding alpha(3)beta(3) subunits leads to hydrolysis of ATP in three separate catalytic sites on the beta subunits.
|
|||||
| Uniprot ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
BTD Probe Info |
 |
C19(0.87) | LDD1700 | [1] | |
|
DBIA Probe Info |
 |
C19(1.01) | LDD0078 | [2] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [3] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [4] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [5] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [4] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [6] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [7] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [8] | |
|
Cinnamaldehyde Probe Info |
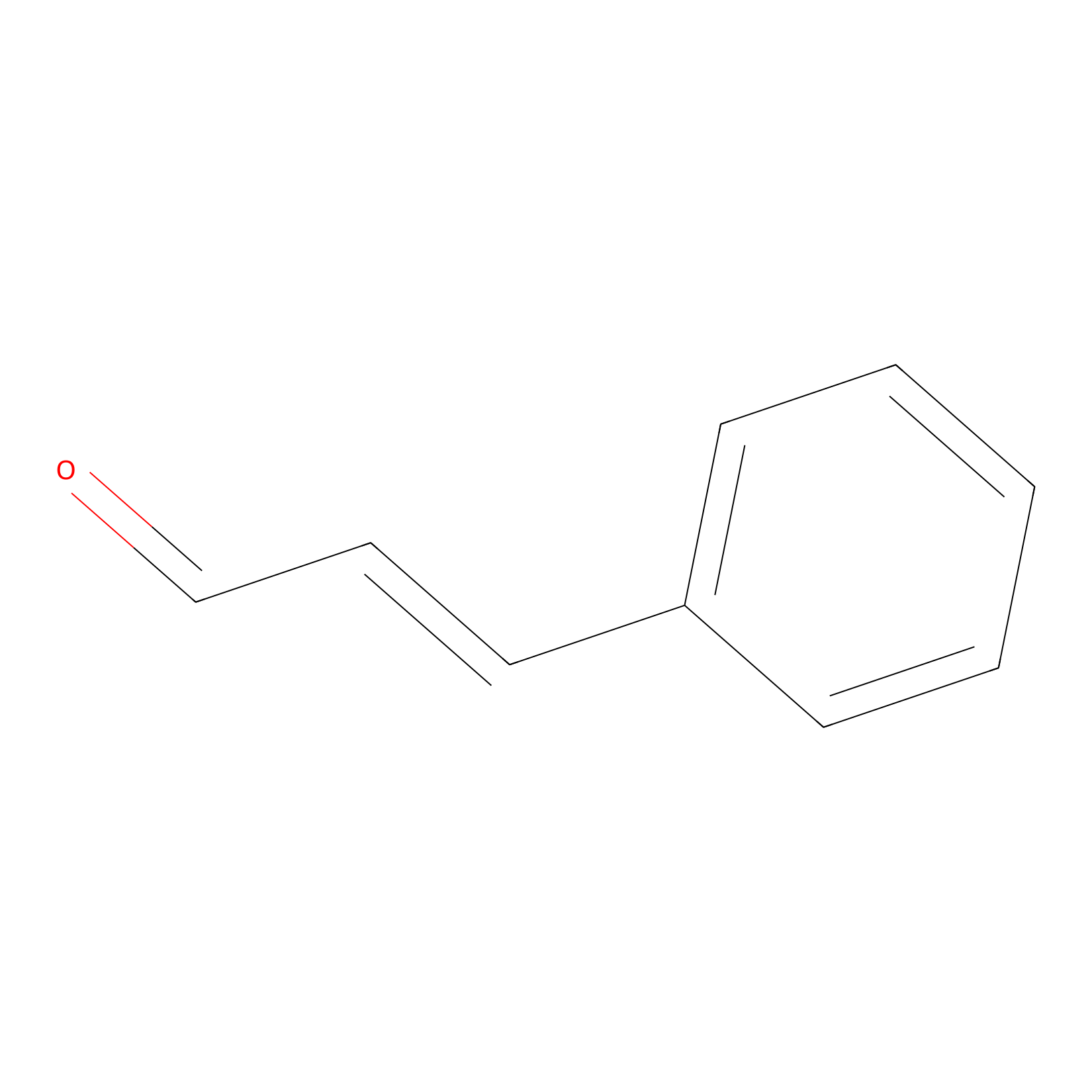 |
N.A. | LDD0220 | [8] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [8] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [8] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [9] | |
|
AOyne Probe Info |
 |
13.00 | LDD0443 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0548 | 1-(4-(Benzo[d][1,3]dioxol-5-ylmethyl)piperazin-1-yl)-2-nitroethan-1-one | MDA-MB-231 | C19(0.53) | LDD2142 | [1] |
| LDCM0519 | 1-(6-methoxy-3,4-dihydroquinolin-1(2H)-yl)-2-nitroethan-1-one | MDA-MB-231 | C19(0.79) | LDD2112 | [1] |
| LDCM0545 | Acetamide | MDA-MB-231 | C19(3.02) | LDD2138 | [1] |
| LDCM0520 | AKOS000195272 | MDA-MB-231 | C19(1.74) | LDD2113 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C19(1.01) | LDD0078 | [2] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [8] |
| LDCM0022 | KB02 | 22RV1 | C19(1.36) | LDD2243 | [11] |
| LDCM0023 | KB03 | 22RV1 | C19(1.77) | LDD2660 | [11] |
| LDCM0024 | KB05 | Hs 936.T | C19(1.68) | LDD3313 | [11] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [8] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C19(1.03) | LDD2093 | [1] |
| LDCM0503 | Nucleophilic fragment 14b | MDA-MB-231 | C19(0.13) | LDD2096 | [1] |
| LDCM0504 | Nucleophilic fragment 15a | MDA-MB-231 | C19(2.95) | LDD2097 | [1] |
| LDCM0507 | Nucleophilic fragment 16b | MDA-MB-231 | C19(0.46) | LDD2100 | [1] |
| LDCM0508 | Nucleophilic fragment 17a | MDA-MB-231 | C19(1.12) | LDD2101 | [1] |
| LDCM0511 | Nucleophilic fragment 18b | MDA-MB-231 | C19(0.54) | LDD2104 | [1] |
| LDCM0515 | Nucleophilic fragment 20b | MDA-MB-231 | C19(0.55) | LDD2108 | [1] |
| LDCM0517 | Nucleophilic fragment 21b | MDA-MB-231 | C19(1.20) | LDD2110 | [1] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C19(1.08) | LDD2111 | [1] |
| LDCM0521 | Nucleophilic fragment 23b | MDA-MB-231 | C19(0.91) | LDD2114 | [1] |
| LDCM0522 | Nucleophilic fragment 24a | MDA-MB-231 | C19(1.27) | LDD2115 | [1] |
| LDCM0525 | Nucleophilic fragment 25b | MDA-MB-231 | C19(0.41) | LDD2118 | [1] |
| LDCM0531 | Nucleophilic fragment 28b | MDA-MB-231 | C19(0.13) | LDD2124 | [1] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C19(5.06) | LDD2125 | [1] |
| LDCM0533 | Nucleophilic fragment 29b | MDA-MB-231 | C19(0.14) | LDD2126 | [1] |
| LDCM0540 | Nucleophilic fragment 35 | MDA-MB-231 | C19(0.92) | LDD2133 | [1] |
| LDCM0541 | Nucleophilic fragment 36 | MDA-MB-231 | C19(1.62) | LDD2134 | [1] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C19(5.68) | LDD2135 | [1] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C19(2.91) | LDD2136 | [1] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C19(0.87) | LDD1700 | [1] |
| LDCM0547 | Nucleophilic fragment 41 | MDA-MB-231 | C19(0.46) | LDD2141 | [1] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C19(1.32) | LDD2143 | [1] |
| LDCM0551 | Nucleophilic fragment 5b | MDA-MB-231 | C19(0.25) | LDD2145 | [1] |
| LDCM0554 | Nucleophilic fragment 7a | MDA-MB-231 | C19(0.69) | LDD2148 | [1] |
References
