Details of the Target
General Information of Target
| Target ID | LDTP13684 | |||||
|---|---|---|---|---|---|---|
| Target Name | NSFL1 cofactor p47 (NSFL1C) | |||||
| Gene Name | NSFL1C | |||||
| Gene ID | 55968 | |||||
| Synonyms |
UBXN2C; NSFL1 cofactor p47; UBX domain-containing protein 2C; p97 cofactor p47 |
|||||
| 3D Structure | ||||||
| Sequence |
MEPPGPVRGPLQDSSWYEPSAELVQTRMAVSLTAAETLALQGTQGQEKMMMMGPKEEEQS
CEYETRLPGNHSTSQEIFRQRFRHLRYQETPGPREALSQLRVLCCEWLRPEKHTKEQILE FLVLEQFLTILPEELQSWVRGHHPKSGEEAVTVLEDLEKGLEPEPQVPGPAHGPAQEEPW EKKESLGAAQEALSIQLQPKETQPFPKSEQVYLHFLSVVTEDGPEPKDKGSLPQPPITEV ESQVFSEKLATDTSTFEATSEGTLELQQRNPKAERLRWSPAQEESFRQMVVIHKEIPTGK KDHECSECGKTFIYNSHLVVHQRVHSGEKPYKCSDCGKTFKQSSNLGQHQRIHTGEKPFE CNECGKAFRWGAHLVQHQRIHSGEKPYECNECGKAFSQSSYLSQHRRIHSGEKPFICKEC GKAYGWCSELIRHRRVHARKEPSH |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
NSFL1C family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Reduces the ATPase activity of VCP. Necessary for the fragmentation of Golgi stacks during mitosis and for VCP-mediated reassembly of Golgi stacks after mitosis. May play a role in VCP-mediated formation of transitional endoplasmic reticulum (tER). Inhibits the activity of CTSL (in vitro). Together with UBXN2B/p37, regulates the centrosomal levels of kinase AURKA/Aurora A during mitotic progression by promoting AURKA removal from centrosomes in prophase. Also, regulates spindle orientation during mitosis.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 1 Probe Info |
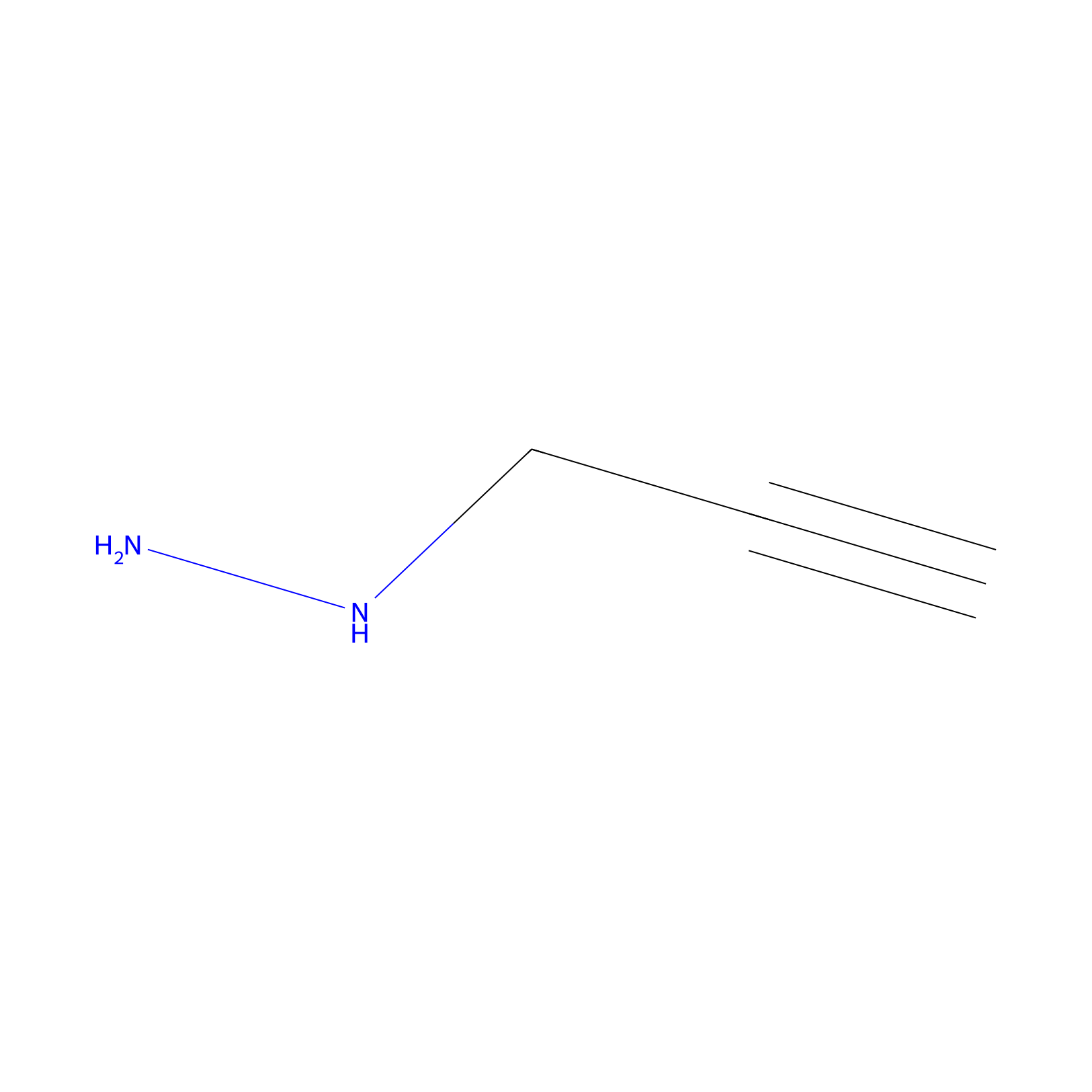 |
8.00 | LDD0387 | [1] | |
|
Probe 1 Probe Info |
 |
Y167(77.61) | LDD3495 | [2] | |
|
HHS-475 Probe Info |
 |
Y167(0.61); Y95(0.99) | LDD0264 | [3] | |
|
HHS-465 Probe Info |
 |
Y155(9.34); Y167(6.08); Y95(4.45) | LDD2237 | [4] | |
|
AMP probe Probe Info |
 |
N.A. | LDD0200 | [5] | |
|
ATP probe Probe Info |
 |
K172(0.00); K124(0.00); K127(0.00); K112(0.00) | LDD0199 | [5] | |
|
m-APA Probe Info |
 |
N.A. | LDD2232 | [6] | |
|
NHS Probe Info |
 |
K172(0.00); K251(0.00) | LDD0010 | [7] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [8] | |
|
STPyne Probe Info |
 |
N.A. | LDD0009 | [7] | |
|
1c-yne Probe Info |
 |
N.A. | LDD0228 | [9] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
HHS-482 Probe Info |
 |
Y155(1.51); Y167(0.96); Y95(1.01) | LDD2239 | [4] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C087 Probe Info |
 |
7.26 | LDD1779 | [11] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [10] |
| LDCM0116 | HHS-0101 | DM93 | Y167(0.61); Y95(0.99) | LDD0264 | [3] |
| LDCM0117 | HHS-0201 | DM93 | Y95(0.87); Y167(0.91) | LDD0265 | [3] |
| LDCM0118 | HHS-0301 | DM93 | Y167(0.65); Y95(0.93) | LDD0266 | [3] |
| LDCM0119 | HHS-0401 | DM93 | Y167(0.49); Y95(0.91) | LDD0267 | [3] |
| LDCM0120 | HHS-0701 | DM93 | Y167(0.69); Y95(0.82) | LDD0268 | [3] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [10] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [10] |
The Interaction Atlas With This Target
References
