Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
AHL-Pu-1 Probe Info |
 |
C791(3.34) | LDD0168 | [1] | |
|
Alkyne-RA190 Probe Info |
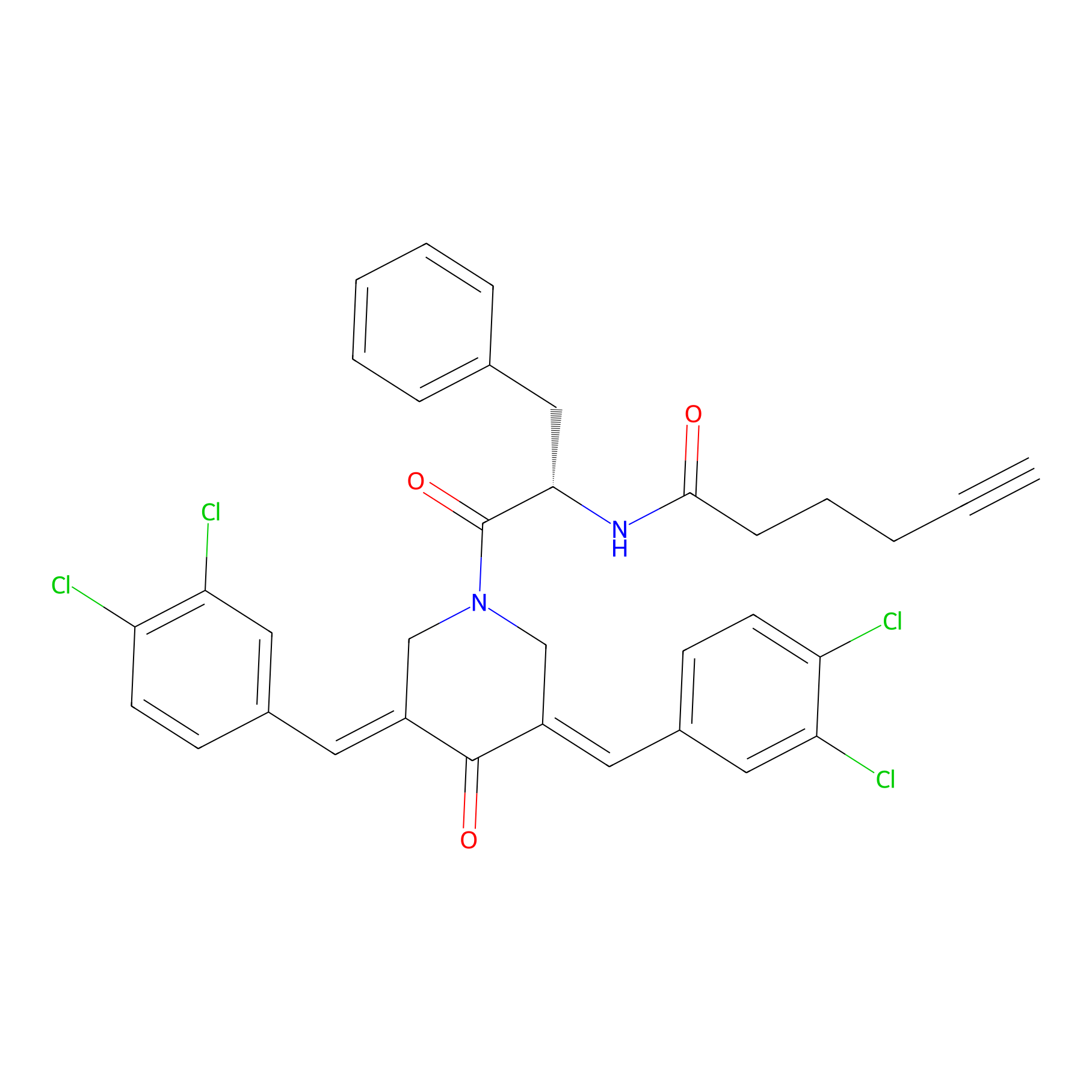 |
2.22 | LDD0299 | [2] | |
|
DBIA Probe Info |
 |
C260(18.11); C150(7.39); C340(5.78); C126(579.08) | LDD0209 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C51(0.00); C487(0.00); C126(0.00); C150(0.00) | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
C51(0.00); C150(0.00); C631(0.00); C126(0.00) | LDD0036 | [4] | |
|
Lodoacetamide azide Probe Info |
 |
C51(0.00); C102(0.00); C126(0.00); C487(0.00) | LDD0037 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C791(3.34) | LDD0168 | [1] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C791(2.56); C51(2.03); C842(2.79) | LDD0169 | [1] |
| LDCM0198 | Dimethyl Fumarate(DMF) | T cell | C150(5.90) | LDD0513 | [5] |
| LDCM0022 | KB02 | T cell-activated | C150(5.51) | LDD1704 | [5] |
| LDCM0023 | KB03 | Jurkat | C260(18.11); C150(7.39); C340(5.78); C126(579.08) | LDD0209 | [3] |
| LDCM0024 | KB05 | G361 | C126(1.47); C340(1.58) | LDD3311 | [6] |
| LDCM0131 | RA190 | MM1.R | 2.22 | LDD0299 | [2] |
References
