Details of the Target
General Information of Target
| Target ID | LDTP07913 | |||||
|---|---|---|---|---|---|---|
| Target Name | GPI inositol-deacylase (PGAP1) | |||||
| Gene Name | PGAP1 | |||||
| Gene ID | 80055 | |||||
| Synonyms |
GPI inositol-deacylase; EC 3.1.-.-; Post-GPI attachment to proteins factor 1; hPGAP1 |
|||||
| 3D Structure | ||||||
| Sequence |
MFLHSVNLWNLAFYVFMVFLATLGLWDVFFGFEENKCSMSYMFEYPEYQKIELPKKLAKR
YPAYELYLYGEGSYAEEHKILPLTGIPVLFLPGNAGSYKQVRSIGSIALRKAEDIDFKYH FDFFSVNFNEELVALYGGSLQKQTKFVHECIKTILKLYKGQEFAPKSVAIIGHSMGGLVA RALLTLKNFKHDLINLLITQATPHVAPVMPLDRFITDFYTTVNNYWILNARHINLTTLSV AGGFRDYQVRSGLTFLPKLSHHTSALSVVSSAVPKTWVSTDHLSIVWCKQLQLTTVRAFF DLIDADTKQITQNSKKKLSVLYHHFIRHPSKHFEENPAIISDLTGTSMWVLVKVSKWTYV AYNESEKIYFTFPLENHRKIYTHVYCQSTMLDTNSWIFACINSTSMCLQGVDLSWKAELL PTIKYLTLRLQDYPSLSHLVVYVPSVRGSKFVVDCEFFKKEKRYIQLPVTHLFSFGLSSR KVVLNTNGLYYNLELLNFGQIYQAFKINVVSKCSAVKEEITSIYRLHIPWSYEDSLTIAQ APSSTEISLKLHIAQPENNTHVALFKMYTSSDCRYEVTVKTSFSQILGQVVRFHGGALPA YVVSNILLAYRGQLYSLFSTGCCLEYATMLDKEAKPYKVDPFVIIIKFLLGYKWFKELWD VLLLPELDAVILTCQSMCFPLISLILFLFGTCTAYWSGLLSSASVRLLSSLWLALKRPSE LPKDIKMISPDLPFLTIVLIIVSWTTCGALAILLSYLYYVFKVVHLQASLTTFKNSQPVN PKHSRRSEKKSNHHKDSSIHHLRLSANDAEDSLRMHSTVINLLTWIVLLSMPSLIYWLKN LRYYFKLNPDPCKPLAFILIPTMAILGNTYTVSIKSSKLLKTTSQFPLPLAVGVIAFGSA HLYRLPCFVFIPLLLHALCNFM |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
GPI inositol-deacylase family
|
|||||
| Subcellular location |
Endoplasmic reticulum membrane
|
|||||
| Function | Involved in inositol deacylation of GPI-anchored proteins. GPI inositol deacylation may important for efficient transport of GPI-anchored proteins from the endoplasmic reticulum to the Golgi. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
HDSF-alk Probe Info |
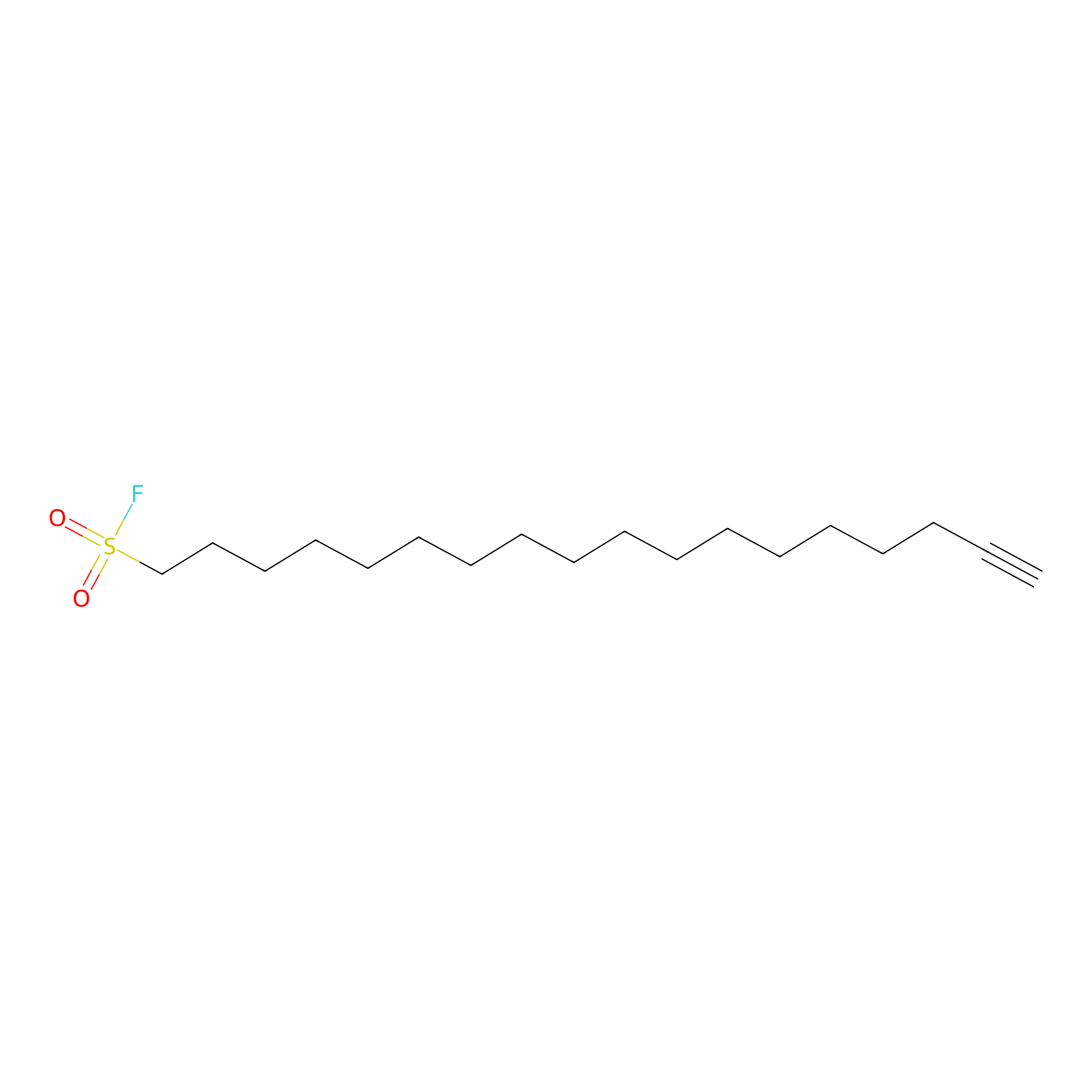 |
2.64 | LDD0197 | [1] | |
|
DBIA Probe Info |
 |
C455(2.52) | LDD3393 | [2] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0214 | AC1 | HEK-293T | C455(1.18) | LDD1507 | [3] |
| LDCM0215 | AC10 | HEK-293T | C455(1.22) | LDD1508 | [3] |
| LDCM0276 | AC17 | HEK-293T | C455(0.91) | LDD1515 | [3] |
| LDCM0277 | AC18 | HEK-293T | C455(1.03) | LDD1516 | [3] |
| LDCM0279 | AC2 | HEK-293T | C455(0.95) | LDD1518 | [3] |
| LDCM0285 | AC25 | HEK-293T | C455(1.33) | LDD1524 | [3] |
| LDCM0286 | AC26 | HEK-293T | C455(0.87) | LDD1525 | [3] |
| LDCM0294 | AC33 | HEK-293T | C455(1.16) | LDD1533 | [3] |
| LDCM0295 | AC34 | HEK-293T | C455(1.28) | LDD1534 | [3] |
| LDCM0303 | AC41 | HEK-293T | C455(1.06) | LDD1542 | [3] |
| LDCM0304 | AC42 | HEK-293T | C455(2.48) | LDD1543 | [3] |
| LDCM0311 | AC49 | HEK-293T | C455(0.99) | LDD1550 | [3] |
| LDCM0313 | AC50 | HEK-293T | C455(1.05) | LDD1552 | [3] |
| LDCM0320 | AC57 | HEK-293T | C455(0.91) | LDD1559 | [3] |
| LDCM0321 | AC58 | HEK-293T | C455(0.90) | LDD1560 | [3] |
| LDCM0356 | AKOS034007680 | HEK-293T | C455(0.98) | LDD1570 | [3] |
| LDCM0369 | CL100 | HEK-293T | C455(1.11) | LDD1573 | [3] |
| LDCM0373 | CL104 | HEK-293T | C455(1.07) | LDD1577 | [3] |
| LDCM0377 | CL108 | HEK-293T | C455(0.99) | LDD1581 | [3] |
| LDCM0382 | CL112 | HEK-293T | C455(0.96) | LDD1586 | [3] |
| LDCM0386 | CL116 | HEK-293T | C455(0.99) | LDD1590 | [3] |
| LDCM0391 | CL120 | HEK-293T | C455(1.25) | LDD1595 | [3] |
| LDCM0395 | CL124 | HEK-293T | C455(1.12) | LDD1599 | [3] |
| LDCM0399 | CL128 | HEK-293T | C455(1.02) | LDD1603 | [3] |
| LDCM0403 | CL16 | HEK-293T | C455(1.02) | LDD1607 | [3] |
| LDCM0404 | CL17 | HEK-293T | C455(1.99) | LDD1608 | [3] |
| LDCM0405 | CL18 | HEK-293T | C455(1.69) | LDD1609 | [3] |
| LDCM0416 | CL28 | HEK-293T | C455(1.33) | LDD1620 | [3] |
| LDCM0417 | CL29 | HEK-293T | C455(1.18) | LDD1621 | [3] |
| LDCM0419 | CL30 | HEK-293T | C455(1.00) | LDD1623 | [3] |
| LDCM0429 | CL4 | HEK-293T | C455(1.32) | LDD1633 | [3] |
| LDCM0430 | CL40 | HEK-293T | C455(1.35) | LDD1634 | [3] |
| LDCM0431 | CL41 | HEK-293T | C455(1.25) | LDD1635 | [3] |
| LDCM0432 | CL42 | HEK-293T | C455(0.96) | LDD1636 | [3] |
| LDCM0440 | CL5 | HEK-293T | C455(1.29) | LDD1644 | [3] |
| LDCM0443 | CL52 | HEK-293T | C455(1.15) | LDD1646 | [3] |
| LDCM0444 | CL53 | HEK-293T | C455(1.68) | LDD1647 | [3] |
| LDCM0445 | CL54 | HEK-293T | C455(0.77) | LDD1648 | [3] |
| LDCM0451 | CL6 | HEK-293T | C455(1.18) | LDD1654 | [3] |
| LDCM0456 | CL64 | HEK-293T | C455(0.92) | LDD1659 | [3] |
| LDCM0457 | CL65 | HEK-293T | C455(1.47) | LDD1660 | [3] |
| LDCM0458 | CL66 | HEK-293T | C455(0.82) | LDD1661 | [3] |
| LDCM0469 | CL76 | HEK-293T | C455(0.89) | LDD1672 | [3] |
| LDCM0470 | CL77 | HEK-293T | C455(1.50) | LDD1673 | [3] |
| LDCM0471 | CL78 | HEK-293T | C455(1.00) | LDD1674 | [3] |
| LDCM0482 | CL88 | HEK-293T | C455(1.53) | LDD1685 | [3] |
| LDCM0483 | CL89 | HEK-293T | C455(0.90) | LDD1686 | [3] |
| LDCM0485 | CL90 | HEK-293T | C455(1.20) | LDD1688 | [3] |
| LDCM0022 | KB02 | A673 | C455(2.91) | LDD2261 | [2] |
| LDCM0023 | KB03 | A673 | C455(4.62) | LDD2678 | [2] |
| LDCM0024 | KB05 | PaTu 8988t | C455(2.52) | LDD3393 | [2] |
References
