Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Alkylaryl probe 2 Probe Info |
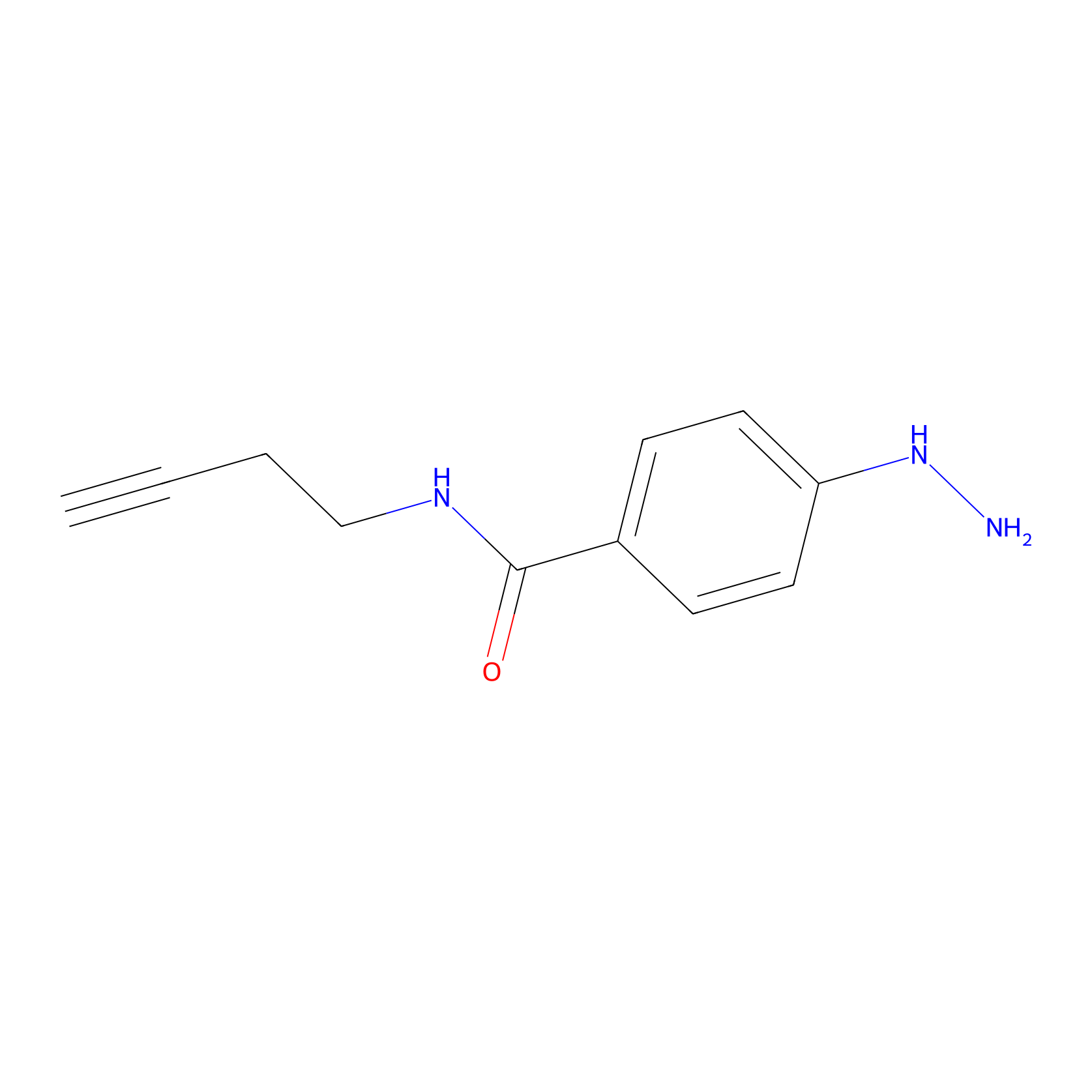 |
3.00 | LDD0390 | [1] | |
|
IPM Probe Info |
 |
C31(4.94) | LDD1701 | [2] | |
|
Acrolein Probe Info |
 |
C268(0.00); C31(0.00) | LDD0221 | [3] | |
|
DBIA Probe Info |
 |
C5(2.76); C8(2.20) | LDD0080 | [4] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0163 | [5] | |
|
NAIA_5 Probe Info |
 |
C31(0.00); C268(0.00); C57(0.00); C466(0.00) | LDD2223 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0226 | AC11 | HEK-293T | C5(1.32) | LDD1509 | [7] |
| LDCM0278 | AC19 | HEK-293T | C5(1.13) | LDD1517 | [7] |
| LDCM0287 | AC27 | HEK-293T | C5(1.31) | LDD1526 | [7] |
| LDCM0290 | AC3 | HEK-293T | C5(1.18) | LDD1529 | [7] |
| LDCM0296 | AC35 | HEK-293T | C5(1.33) | LDD1535 | [7] |
| LDCM0305 | AC43 | HEK-293T | C5(1.32) | LDD1544 | [7] |
| LDCM0314 | AC51 | HEK-293T | C5(1.46) | LDD1553 | [7] |
| LDCM0322 | AC59 | HEK-293T | C5(1.30) | LDD1561 | [7] |
| LDCM0108 | Chloroacetamide | HeLa | C268(0.00); C31(0.00); C57(0.00) | LDD0222 | [3] |
| LDCM0632 | CL-Sc | Hep-G2 | C5(0.57) | LDD2227 | [6] |
| LDCM0406 | CL19 | HEK-293T | C5(1.29) | LDD1610 | [7] |
| LDCM0420 | CL31 | HEK-293T | C5(1.39) | LDD1624 | [7] |
| LDCM0433 | CL43 | HEK-293T | C5(1.24) | LDD1637 | [7] |
| LDCM0446 | CL55 | HEK-293T | C5(1.36) | LDD1649 | [7] |
| LDCM0459 | CL67 | HEK-293T | C5(1.22) | LDD1662 | [7] |
| LDCM0462 | CL7 | HEK-293T | C5(1.26) | LDD1665 | [7] |
| LDCM0472 | CL79 | HEK-293T | C5(1.24) | LDD1675 | [7] |
| LDCM0486 | CL91 | HEK-293T | C5(1.19) | LDD1689 | [7] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C31(2.04) | LDD1702 | [2] |
| LDCM0107 | IAA | HeLa | C268(0.00); C31(0.00) | LDD0221 | [3] |
| LDCM0022 | KB02 | HCT 116 | C5(2.76); C8(2.20) | LDD0080 | [4] |
| LDCM0023 | KB03 | HCT 116 | C5(1.11); C8(1.17) | LDD0081 | [4] |
| LDCM0024 | KB05 | HCT 116 | C5(2.09); C8(1.73) | LDD0082 | [4] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0224 | [3] |
| LDCM0099 | Phenelzine | HEK-293T | 3.00 | LDD0390 | [1] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| 52 kDa repressor of the inhibitor of the protein kinase (THAP12) | . | O43422 | |||
References
