Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STPyne Probe Info |
 |
K66(2.33) | LDD0277 | [1] | |
|
DBIA Probe Info |
 |
C241(2.31) | LDD3314 | [2] | |
|
Curcusone 37 Probe Info |
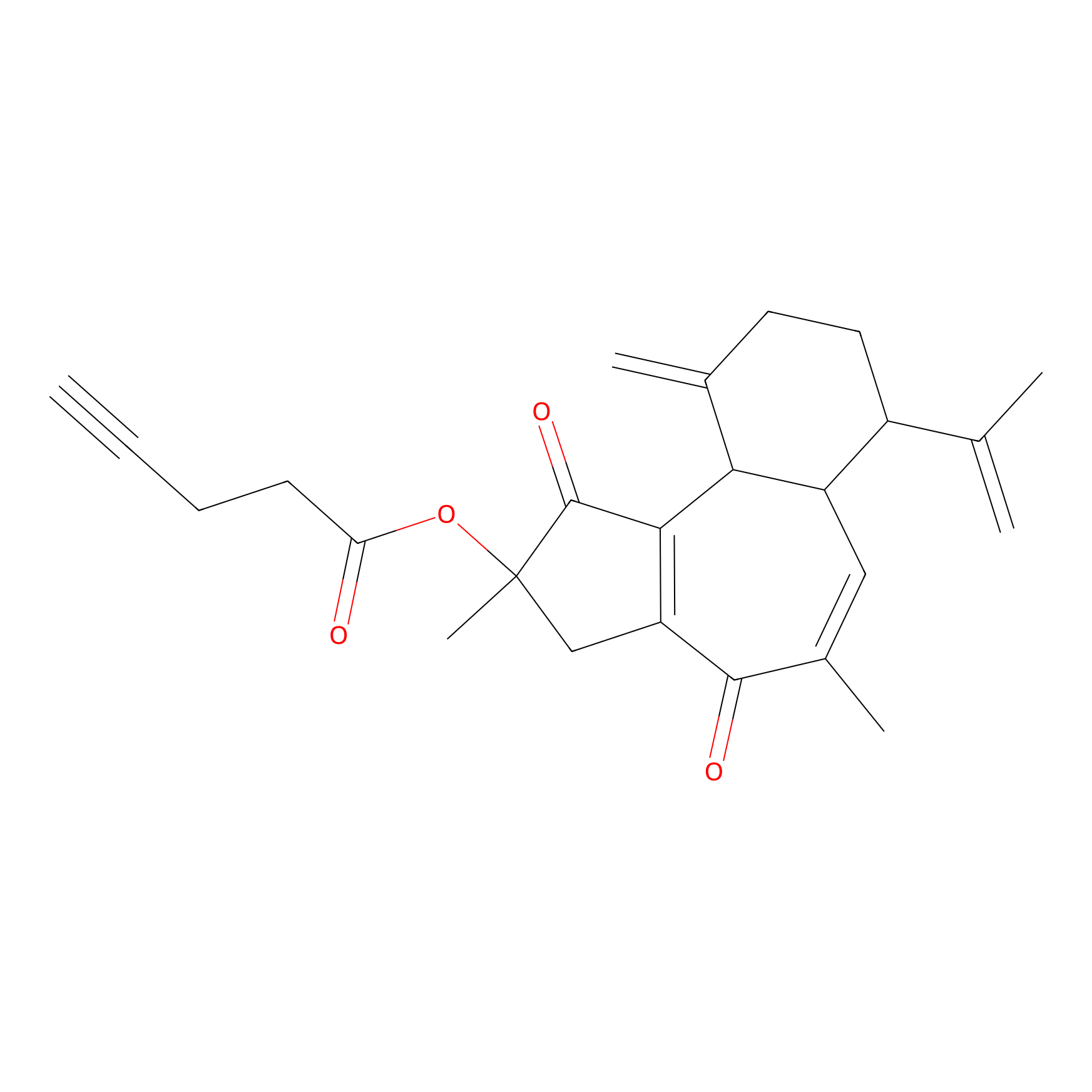 |
2.63 | LDD0188 | [3] | |
|
IA-alkyne Probe Info |
 |
C319(20.00) | LDD0513 | [4] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2225 | [5] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0033 | Curcusone1d | MCF-7 | 2.63 | LDD0188 | [3] |
| LDCM0198 | Dimethyl Fumarate(DMF) | T cell | C319(20.00) | LDD0513 | [4] |
| LDCM0022 | KB02 | T cell | C319(20.00) | LDD1703 | [4] |
| LDCM0023 | KB03 | 8305C | C241(2.89) | LDD2665 | [2] |
| LDCM0024 | KB05 | IGR37 | C241(2.31) | LDD3314 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Signal transducer and activator of transcription 2 (STAT2) | Transcription factor STAT family | P52630 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Interferon alpha/beta receptor 2 (IFNAR2) | Type II cytokine receptor family | P48551 | |||
References
