Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-1 Probe Info |
 |
100.00 | LDD0444 | [1] | |
|
HHS-475 Probe Info |
 |
Y241(0.75); Y54(0.77); Y70(0.92); Y189(1.75) | LDD0264 | [2] | |
|
HHS-465 Probe Info |
 |
Y189(10.00); Y241(10.00); Y54(10.00); Y70(9.20) | LDD2237 | [3] | |
|
ATP probe Probe Info |
 |
K62(0.00); K329(0.00); K51(0.00) | LDD0035 | [4] | |
|
AZ-9 Probe Info |
 |
D26(0.00); E242(0.00) | LDD0395 | [5] | |
|
NHS Probe Info |
 |
K292(0.00); K51(0.00); K327(0.00); K69(0.00) | LDD0010 | [6] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Diazir Probe Info |
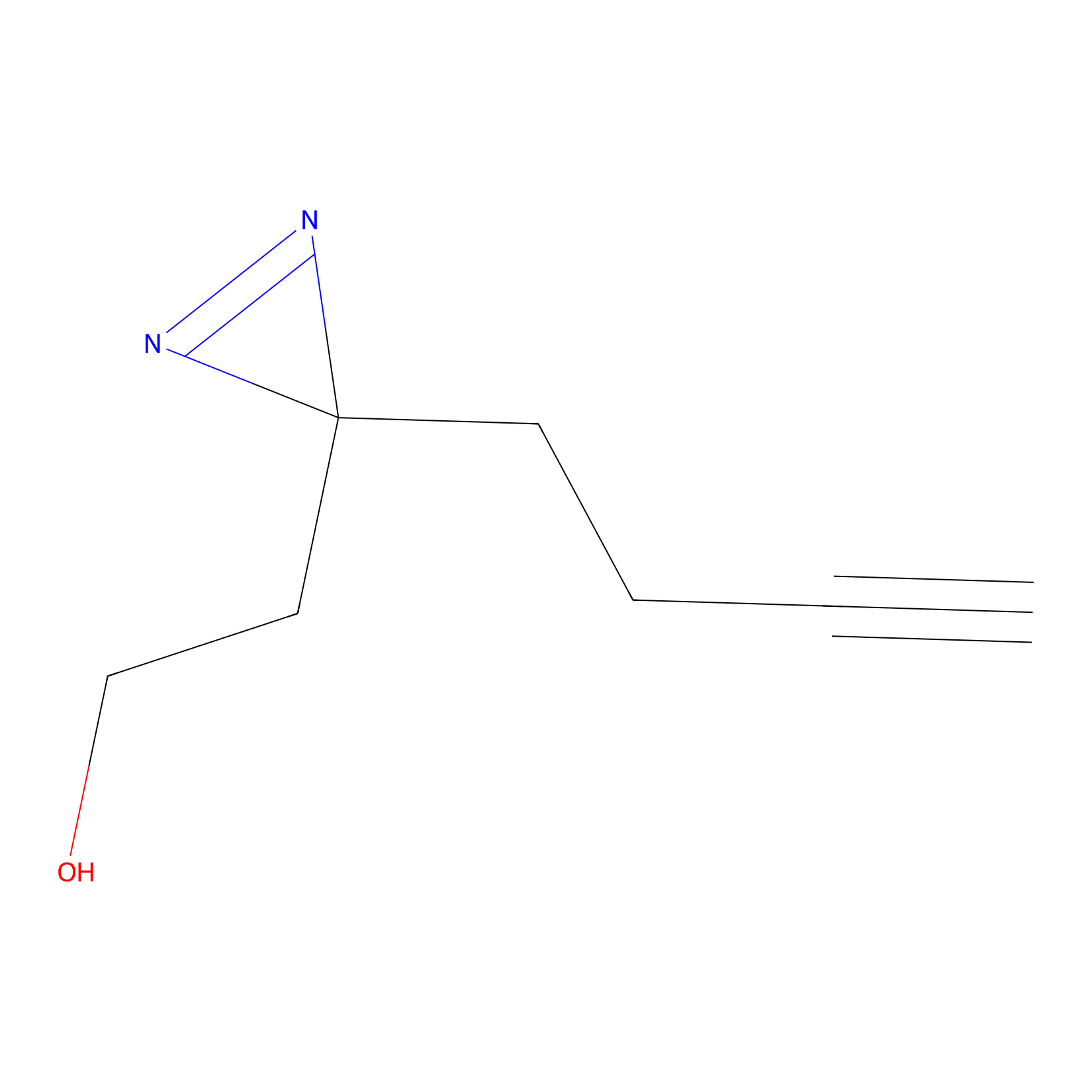 |
E242(0.00); E254(0.00); E84(0.00); Y241(0.00) | LDD0011 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0116 | HHS-0101 | DM93 | Y241(0.75); Y54(0.77); Y70(0.92); Y189(1.75) | LDD0264 | [2] |
| LDCM0117 | HHS-0201 | DM93 | Y241(0.62); Y54(0.67); Y70(0.87); Y189(1.25) | LDD0265 | [2] |
| LDCM0118 | HHS-0301 | DM93 | Y54(0.71); Y70(0.85); Y189(1.11); Y241(1.16) | LDD0266 | [2] |
| LDCM0119 | HHS-0401 | DM93 | Y241(0.73); Y54(0.75); Y70(0.97); Y189(1.59) | LDD0267 | [2] |
| LDCM0120 | HHS-0701 | DM93 | Y241(0.52); Y54(0.71); Y70(0.72); Y189(1.00) | LDD0268 | [2] |
References
