Details of the Target
General Information of Target
| Target ID | LDTP04873 | |||||
|---|---|---|---|---|---|---|
| Target Name | Reactive oxygen species modulator 1 (ROMO1) | |||||
| Gene Name | ROMO1 | |||||
| Gene ID | 140823 | |||||
| Synonyms |
C20orf52; Reactive oxygen species modulator 1; ROS modulator 1; Epididymis tissue protein Li 175; Glyrichin; Mitochondrial targeting GxxxG motif protein; MTGM; Protein MGR2 homolog |
|||||
| 3D Structure | ||||||
| Sequence |
MPVAVGPYGQSQPSCFDRVKMGFVMGCAVGMAAGALFGTFSCLRIGMRGRELMGGIGKTM
MQSGGTFGTFMAIGMGIRC |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
MGR2 family
|
|||||
| Subcellular location |
Mitochondrion inner membrane
|
|||||
| Function |
Induces production of reactive oxygen species (ROS) which are necessary for cell proliferation. May play a role in inducing oxidative DNA damage and replicative senescence. May play a role in the coordination of mitochondrial morphology and cell proliferation.; Has antibacterial activity against a variety of bacteria including S.aureus, P.aeruginosa and M.tuberculosis. Acts by inducing bacterial membrane breakage.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [1] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [2] | |
|
BTD Probe Info |
 |
C15(1.17) | LDD2090 | [3] | |
|
ART-yne Probe Info |
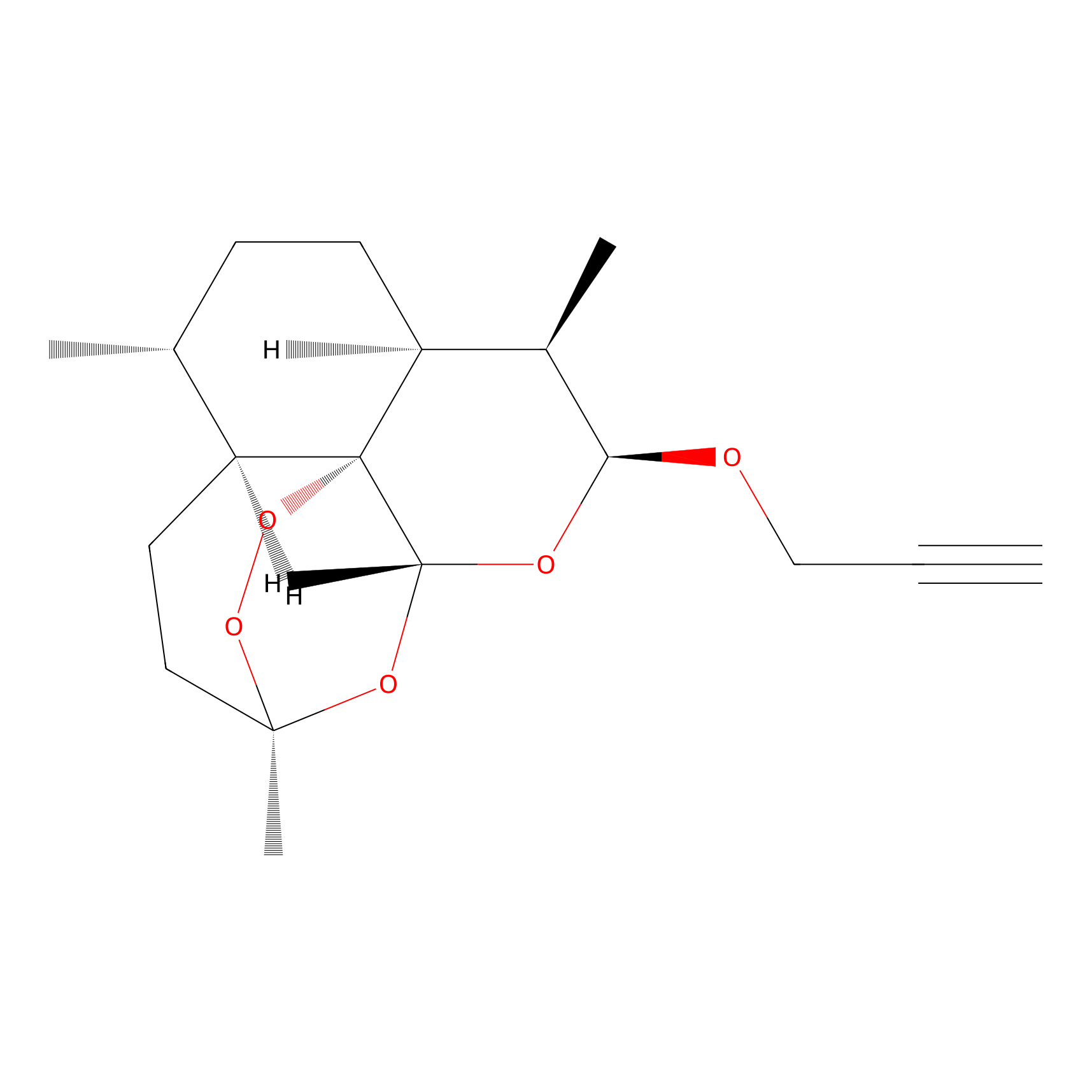 |
2.12 | LDD0234 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
N.A. | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0032 | [6] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [5] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [7] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [7] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [8] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [9] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C15(0.88) | LDD2117 | [3] |
| LDCM0091 | Astemisinin | MCF-7 | 2.12 | LDD0234 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [9] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C15(6.59) | LDD1702 | [3] |
| LDCM0625 | F8 | Ramos | C15(0.83) | LDD2187 | [11] |
| LDCM0572 | Fragment10 | Ramos | C15(0.85) | LDD2189 | [11] |
| LDCM0574 | Fragment12 | Ramos | C15(1.08) | LDD2191 | [11] |
| LDCM0575 | Fragment13 | Ramos | C15(0.95) | LDD2192 | [11] |
| LDCM0576 | Fragment14 | Ramos | C15(4.77) | LDD2193 | [11] |
| LDCM0579 | Fragment20 | Ramos | C15(0.97) | LDD2194 | [11] |
| LDCM0580 | Fragment21 | Ramos | C15(1.02) | LDD2195 | [11] |
| LDCM0582 | Fragment23 | Ramos | C15(0.67) | LDD2196 | [11] |
| LDCM0578 | Fragment27 | Ramos | C15(0.66) | LDD2197 | [11] |
| LDCM0586 | Fragment28 | Ramos | C15(0.88) | LDD2198 | [11] |
| LDCM0588 | Fragment30 | Ramos | C15(0.99) | LDD2199 | [11] |
| LDCM0589 | Fragment31 | Ramos | C15(1.79) | LDD2200 | [11] |
| LDCM0590 | Fragment32 | Ramos | C15(0.86) | LDD2201 | [11] |
| LDCM0468 | Fragment33 | Ramos | C15(1.00) | LDD2202 | [11] |
| LDCM0596 | Fragment38 | Ramos | C15(0.98) | LDD2203 | [11] |
| LDCM0566 | Fragment4 | Ramos | C15(1.40) | LDD2184 | [11] |
| LDCM0610 | Fragment52 | Ramos | C15(1.23) | LDD2204 | [11] |
| LDCM0614 | Fragment56 | Ramos | C15(0.92) | LDD2205 | [11] |
| LDCM0569 | Fragment7 | Ramos | C15(0.85) | LDD2186 | [11] |
| LDCM0571 | Fragment9 | Ramos | C15(1.07) | LDD2188 | [11] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [9] |
| LDCM0022 | KB02 | Ramos | C15(0.94) | LDD2182 | [11] |
| LDCM0023 | KB03 | MDA-MB-231 | C15(4.27) | LDD1701 | [3] |
| LDCM0024 | KB05 | Ramos | C15(1.43) | LDD2185 | [11] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0229 | [9] |
| LDCM0497 | Nucleophilic fragment 11b | MDA-MB-231 | C15(1.17) | LDD2090 | [3] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C15(1.01) | LDD2129 | [3] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C15(0.33) | LDD2207 | [12] |
The Interaction Atlas With This Target
References
