Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
OPA-S-S-alkyne Probe Info |
 |
K109(2.61) | LDD3494 | [1] | |
|
Alkyne-RA190 Probe Info |
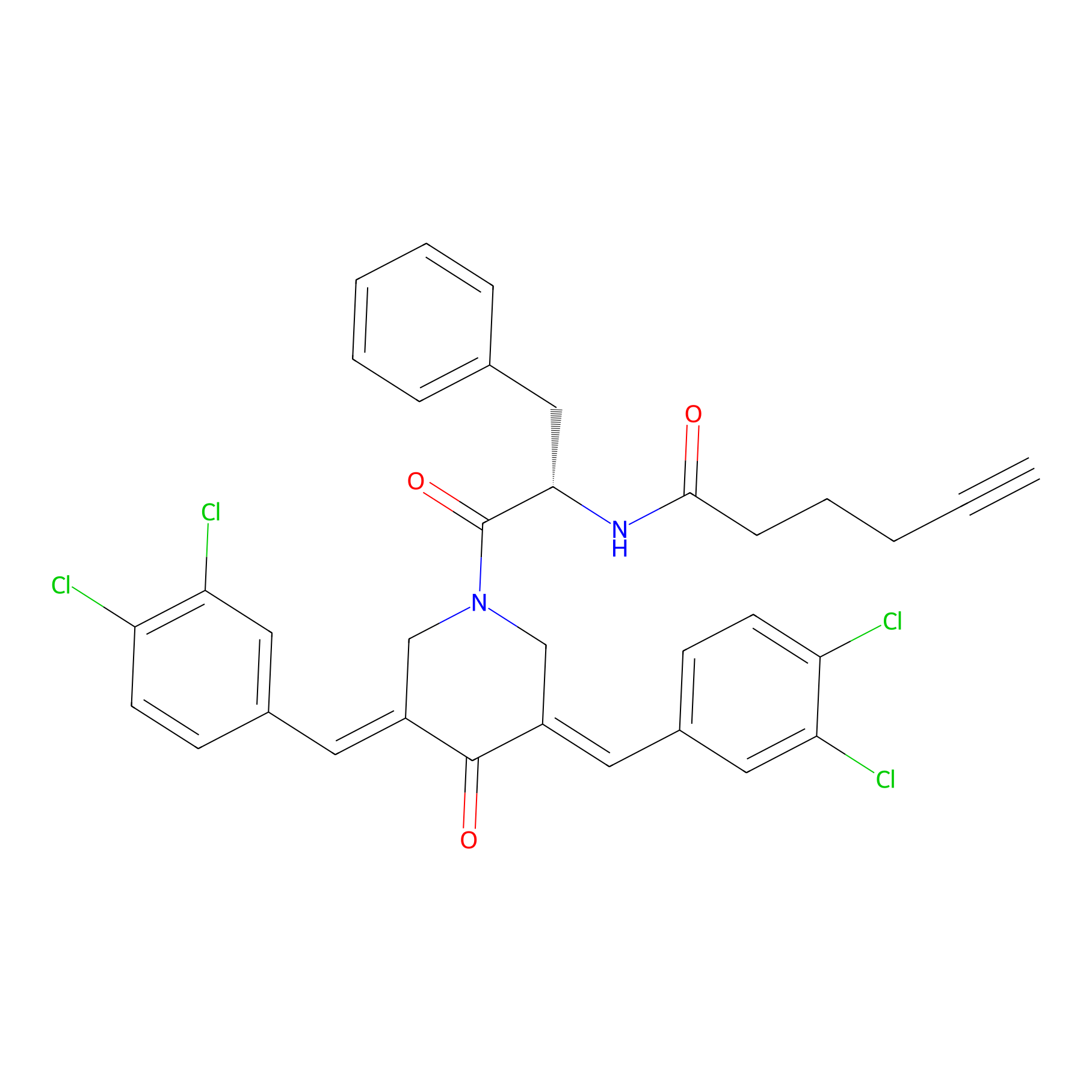 |
2.07 | LDD0303 | [2] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
C153 Probe Info |
 |
21.56 | LDD1834 | [3] | |
|
FFF probe11 Probe Info |
 |
20.00 | LDD0471 | [4] | |
|
FFF probe13 Probe Info |
 |
20.00 | LDD0475 | [4] | |
|
FFF probe14 Probe Info |
 |
20.00 | LDD0477 | [4] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Basigin (BSG) | . | P35613 | |||
| Triggering receptor expressed on myeloid cells 2 (TREM2) | . | Q9NZC2 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Tumor necrosis factor receptor superfamily member 11B (TNFRSF11B) | . | O00300 | |||
Other
The Drug(s) Related To This Target
Phase 2
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| R-flurbiprofen | Small molecular drug | D05FGR | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| (S)-flurbiprofen | Small molecular drug | D0G2IC | |||
Investigative
References
