Details of the Target
General Information of Target
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| CHL1 | SNV: p.A409S | DBIA Probe Info | |||
| HSC3 | SNV: p.D103H | . | |||
| KELLY | SNV: p.A282D | DBIA Probe Info | |||
| NUGC3 | SNV: p.G177Ter | . | |||
| SW1271 | SNV: p.D277G | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
15.00 | LDD0402 | [1] | |
|
TH211 Probe Info |
 |
Y259(9.81) | LDD0257 | [2] | |
|
DBIA Probe Info |
 |
C175(1.47) | LDD3325 | [3] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C137(0.00); C245(0.00); C41(0.00) | LDD0038 | [4] | |
|
IA-alkyne Probe Info |
 |
C245(0.00); C41(0.00) | LDD0036 | [4] | |
|
IPIAA_L Probe Info |
 |
N.A. | LDD0031 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
C137(0.00); C245(0.00); C41(0.00) | LDD0037 | [4] | |
|
IPM Probe Info |
 |
N.A. | LDD2156 | [6] | |
|
W1 Probe Info |
 |
N.A. | LDD0236 | [7] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [8] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [9] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DFG-out-4 Probe Info |
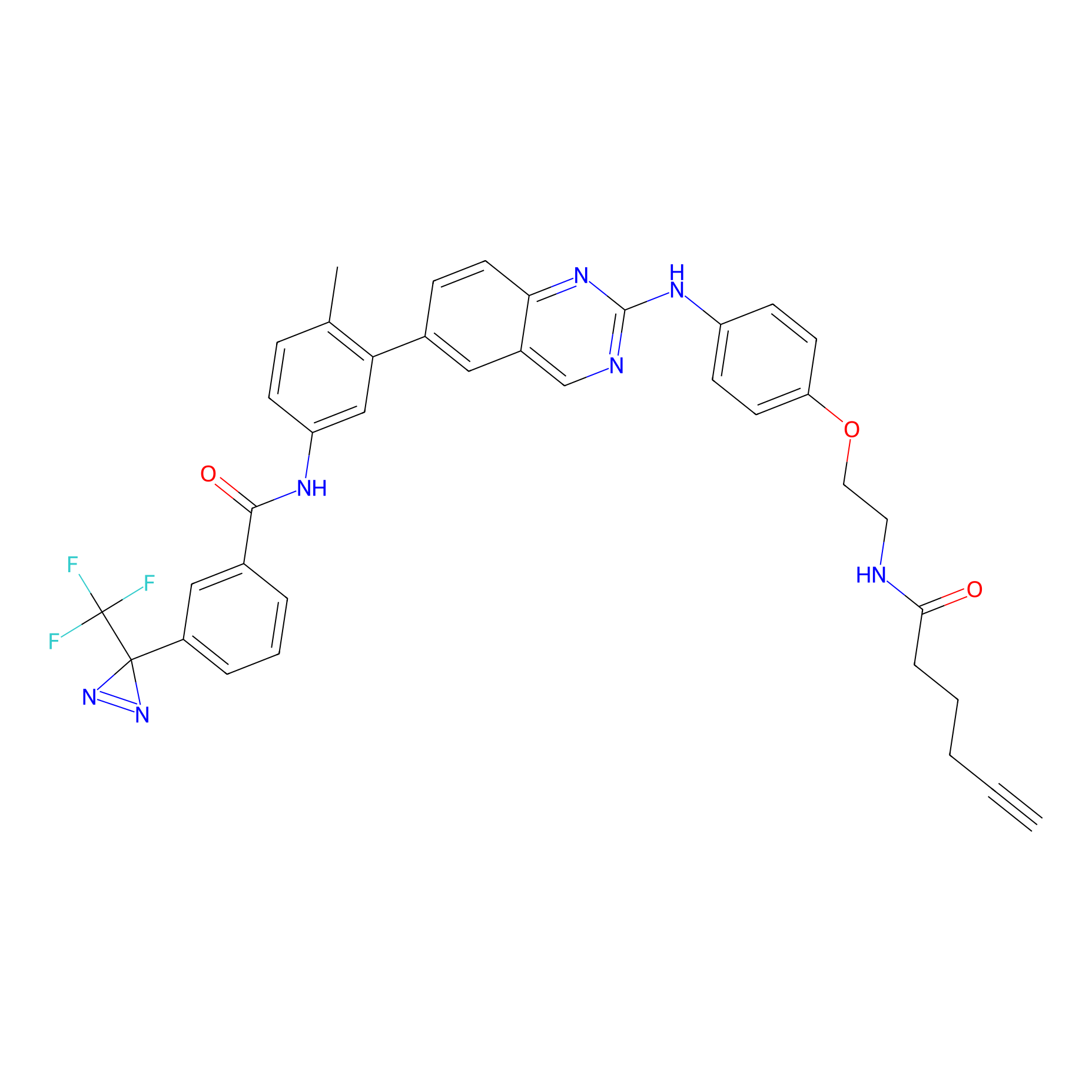 |
6.30 | LDD0075 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 12.20 | LDD0403 | [1] |
| LDCM0632 | CL-Sc | Hep-G2 | C245(1.26) | LDD2227 | [9] |
| LDCM0017 | DFG-out-2 | A431 | 6.30 | LDD0075 | [10] |
| LDCM0625 | F8 | Ramos | C245(1.36) | LDD2187 | [11] |
| LDCM0573 | Fragment11 | Ramos | C245(2.27) | LDD2190 | [11] |
| LDCM0575 | Fragment13 | Ramos | C245(1.01) | LDD2192 | [11] |
| LDCM0576 | Fragment14 | Ramos | C245(0.89) | LDD2193 | [11] |
| LDCM0580 | Fragment21 | Ramos | C245(0.87) | LDD2195 | [11] |
| LDCM0582 | Fragment23 | Ramos | C245(8.50) | LDD2196 | [11] |
| LDCM0578 | Fragment27 | Ramos | C245(0.41) | LDD2197 | [11] |
| LDCM0586 | Fragment28 | Ramos | C245(0.89) | LDD2198 | [11] |
| LDCM0588 | Fragment30 | Ramos | C245(0.82) | LDD2199 | [11] |
| LDCM0589 | Fragment31 | Ramos | C245(0.82) | LDD2200 | [11] |
| LDCM0468 | Fragment33 | Ramos | C245(0.78) | LDD2202 | [11] |
| LDCM0596 | Fragment38 | Ramos | C245(0.60) | LDD2203 | [11] |
| LDCM0566 | Fragment4 | Ramos | C245(1.56) | LDD2184 | [11] |
| LDCM0614 | Fragment56 | Ramos | C245(0.60) | LDD2205 | [11] |
| LDCM0022 | KB02 | Ramos | C245(2.54) | LDD2182 | [11] |
| LDCM0023 | KB03 | 42-MG-BA | C283(1.97) | LDD2661 | [3] |
| LDCM0024 | KB05 | UACC257 | C175(1.47) | LDD3325 | [3] |
| LDCM0112 | W16 | Hep-G2 | C245(1.20) | LDD0239 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor Jun (JUN) | BZIP family | P05412 | |||
Other
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Minocycline | Small molecular drug | DB01017 | |||
| Tamoxifen | Small molecular drug | DB00675 | |||
Phase 1
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Nkp-1339 | Small molecular drug | D08SIT | |||
Preclinical
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Cor-d | Small molecular drug | DOJ6D9 | |||
Investigative
Patented
References
