Details of the Target
General Information of Target
| Target ID | LDTP02972 | |||||
|---|---|---|---|---|---|---|
| Target Name | ATP synthase-coupling factor 6, mitochondrial (ATP5PF) | |||||
| Gene Name | ATP5PF | |||||
| Gene ID | 522 | |||||
| Synonyms |
ATP5A; ATP5J; ATPM; ATP synthase-coupling factor 6, mitochondrial; ATPase subunit F6; ATP synthase peripheral stalk subunit F6 |
|||||
| 3D Structure | ||||||
| Sequence |
MILQRLFRFSSVIRSAVSVHLRRNIGVTAVAFNKELDPIQKLFVDKIREYKSKRQTSGGP
VDASSEYQQELERELFKLKQMFGNADMNTFPTFKFEDPKFEVIEKPQA |
|||||
| Target Bioclass |
Transporter and channel
|
|||||
| Family |
Eukaryotic ATPase subunit F6 family
|
|||||
| Subcellular location |
Mitochondrion
|
|||||
| Function |
Mitochondrial membrane ATP synthase (F(1)F(0) ATP synthase or Complex V) produces ATP from ADP in the presence of a proton gradient across the membrane which is generated by electron transport complexes of the respiratory chain. F-type ATPases consist of two structural domains, F(1) - containing the extramembraneous catalytic core and F(0) - containing the membrane proton channel, linked together by a central stalk and a peripheral stalk. During catalysis, ATP synthesis in the catalytic domain of F(1) is coupled via a rotary mechanism of the central stalk subunits to proton translocation. Part of the complex F(0) domain and the peripheric stalk, which acts as a stator to hold the catalytic alpha(3)beta(3) subcomplex and subunit a/ATP6 static relative to the rotary elements. Also involved in the restoration of oligomycin-sensitive ATPase activity to depleted F1-F0 complexes.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K46(1.00) | LDD0274 | [1] | |
|
STPyne Probe Info |
 |
K105(1.00); K34(6.67); K46(5.23); K79(7.20) | LDD0277 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K105(3.04); K46(4.70) | LDD3494 | [2] | |
|
Probe 1 Probe Info |
 |
Y67(14.24) | LDD3495 | [3] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [4] | |
|
1c-yne Probe Info |
 |
K99(0.00); K79(0.00) | LDD0228 | [5] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [5] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
Staurosporine capture compound Probe Info |
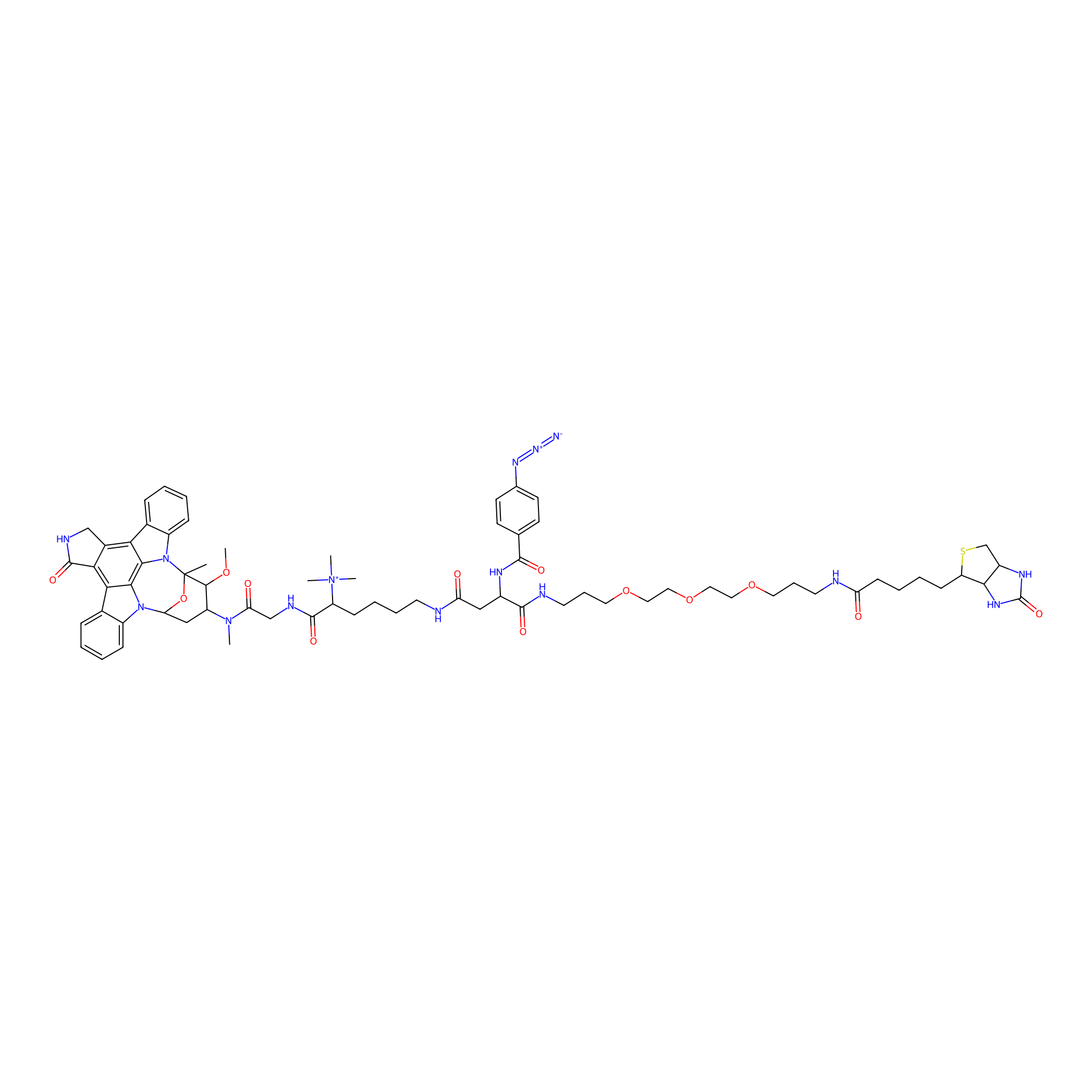 |
N.A. | LDD0083 | [6] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Sialic acid-binding Ig-like lectin 12 (SIGLEC12) | SIGLEC (sialic acid binding Ig-like lectin) family | Q96PQ1 | |||
Other
References
