Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
YN-4 Probe Info |
 |
100.00 | LDD0445 | [1] | |
|
IPM Probe Info |
 |
N.A. | LDD0241 | [2] | |
|
DBIA Probe Info |
 |
C45(1.45) | LDD3354 | [3] | |
|
BTD Probe Info |
 |
C187(2.92) | LDD1700 | [4] | |
|
IA-alkyne Probe Info |
 |
C45(0.00); C187(0.00); C200(0.00) | LDD0162 | [5] | |
|
WYneC Probe Info |
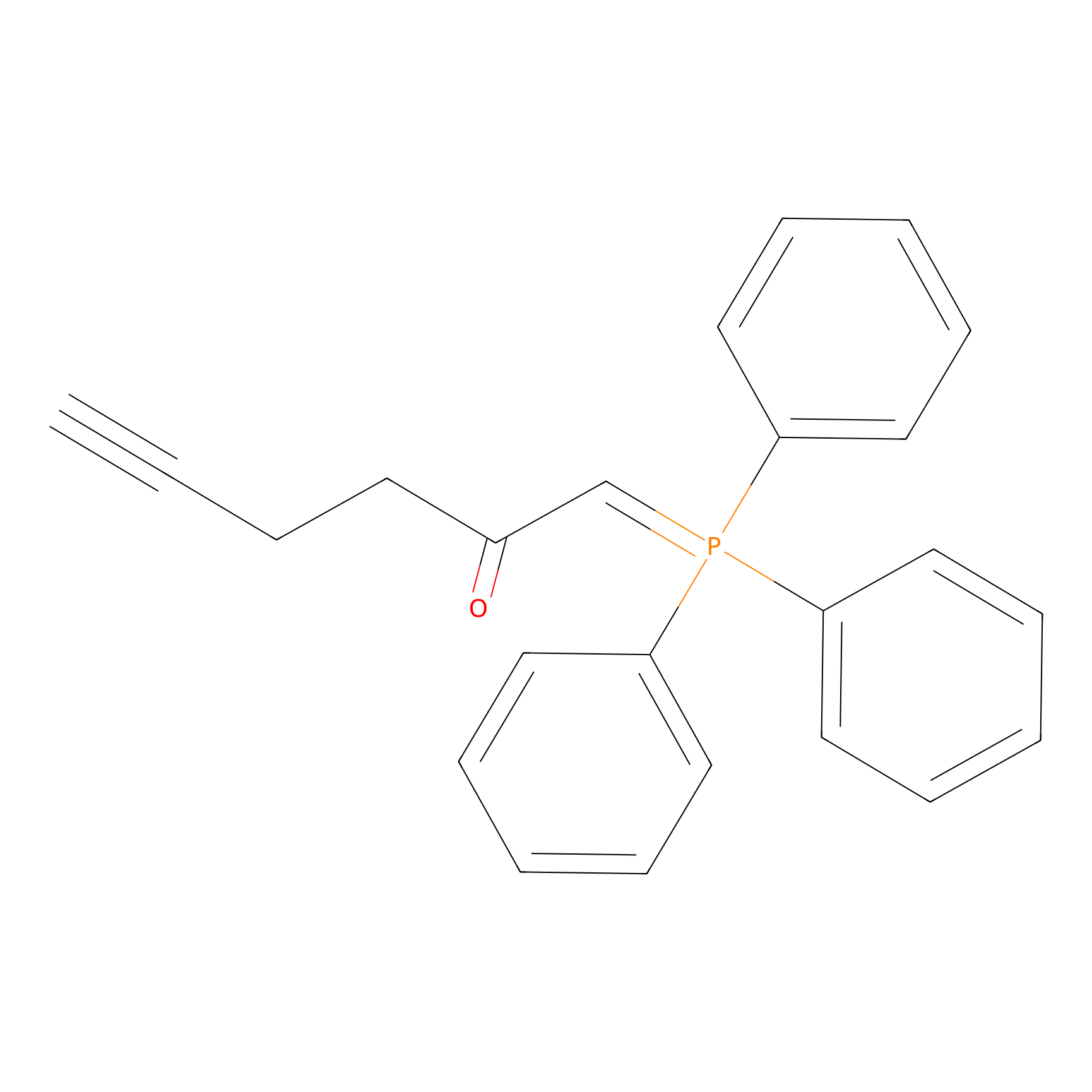 |
N.A. | LDD0014 | [6] | |
|
WYneN Probe Info |
 |
C299(0.00); C187(0.00); C45(0.00) | LDD0021 | [6] | |
|
WYneO Probe Info |
 |
C299(0.00); C187(0.00) | LDD0022 | [6] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [7] | |
|
HHS-475 Probe Info |
 |
Y190(2.32); Y199(1.47); Y47(1.50); Y49(1.60) | LDD2238 | [8] | |
|
HHS-482 Probe Info |
 |
Y178(1.16); Y190(1.07); Y210(1.35); Y47(1.12) | LDD2239 | [8] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0022 | KB02 | A2780 | C45(1.49) | LDD2254 | [3] |
| LDCM0023 | KB03 | A2780 | C45(1.52) | LDD2671 | [3] |
| LDCM0024 | KB05 | NCI-H2122 | C45(1.45) | LDD3354 | [3] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C187(1.04); C200(1.05) | LDD2093 | [4] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C187(1.36); C200(1.04) | LDD2099 | [4] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C187(0.67); C200(0.45) | LDD2109 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C187(1.15); C200(0.94) | LDD2111 | [4] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C187(1.12) | LDD2119 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C187(0.98) | LDD2129 | [4] |
| LDCM0543 | Nucleophilic fragment 38 | MDA-MB-231 | C187(1.64) | LDD2136 | [4] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C187(2.92) | LDD1700 | [4] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Acetyl-CoA carboxylase 1 (ACACA) | . | Q13085 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Telomeric repeat-binding factor 2-interacting protein 1 (TERF2IP) | RAP1 family | Q9NYB0 | |||
The Drug(s) Related To This Target
Approved
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Lumateperone | Small molecular drug | DB06077 | |||
| Sulindac | Small molecular drug | DB00605 | |||
Investigative
Discontinued
| Drug Name | Drug Type | External ID | |||
|---|---|---|---|---|---|
| Tolrestat | Small molecular drug | DB02383 | |||
References
