Details of the Target
General Information of Target
| Target ID | LDTP12923 | |||||
|---|---|---|---|---|---|---|
| Target Name | EH domain-containing protein 2 (EHD2) | |||||
| Gene Name | EHD2 | |||||
| Gene ID | 30846 | |||||
| Synonyms |
PAST2; EH domain-containing protein 2; PAST homolog 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MFSWLGTDDRRRKDPEVFQTVSEGLKKLYKSKLLPLEEHYRFHEFHSPALEDADFDNKPM
VLLVGQYSTGKTTFIRYLLEQDFPGMRIGPEPTTDSFIAVMQGDMEGIIPGNALVVDPKK PFRKLNAFGNAFLNRFVCAQLPNPVLESISVIDTPGILSGEKQRISRGYDFAAVLEWFAE RVDRIILLFDAHKLDISDEFSEVIKALKNHEDKMRVVLNKADQIETQQLMRVYGALMWSL GKIVNTPEVIRVYIGSFWSHPLLIPDNRKLFEAEEQDLFRDIQSLPRNAALRKLNDLIKR ARLAKVHAYIISSLKKEMPSVFGKDNKKKELVNNLAEIYGRIEREHQISPGDFPNLKRMQ DQLQAQDFSKFQPLKSKLLEVVDDMLAHDIAQLMVLVRQEESQRPIQMVKGGAFEGTLHG PFGHGYGEGAGEGIDDAEWVVARDKPMYDEIFYTLSPVDGKITGANAKKEMVRSKLPNSV LGKIWKLADIDKDGMLDDDEFALANHLIKVKLEGHELPNELPAHLLPPSKRKVAE |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
TRAFAC class dynamin-like GTPase superfamily, Dynamin/Fzo/YdjA family, EHD subfamily
|
|||||
| Subcellular location |
Cell membrane
|
|||||
| Function |
ATP- and membrane-binding protein that controls membrane reorganization/tubulation upon ATP hydrolysis. Plays a role in membrane trafficking between the plasma membrane and endosomes. Important for the internalization of GLUT4. Required for fusion of myoblasts to skeletal muscle myotubes. Required for normal translocation of FER1L5 to the plasma membrane. Regulates the equilibrium between cell surface-associated and cell surface-dissociated caveolae by constraining caveolae at the cell membrane.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
P2 Probe Info |
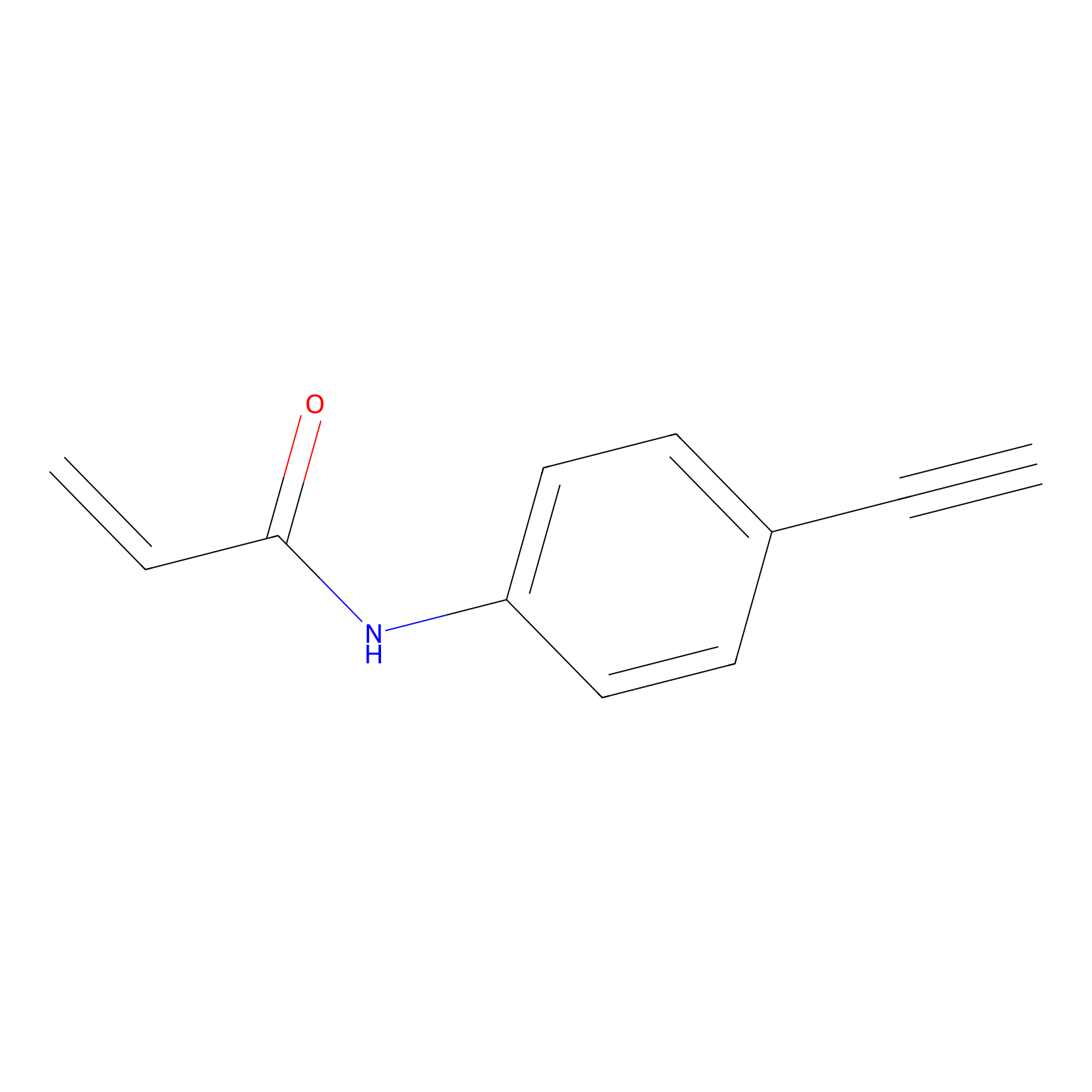 |
4.08 | LDD0449 | [1] | |
|
ONAyne Probe Info |
 |
K213(0.93); K32(0.75) | LDD0274 | [2] | |
|
STPyne Probe Info |
 |
K124(8.71); K213(10.00); K220(6.46); K299(6.67) | LDD0277 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K491(1.25); K220(2.14); K32(2.92) | LDD3494 | [3] | |
|
BTD Probe Info |
 |
C356(0.73) | LDD2107 | [4] | |
|
5E-2FA Probe Info |
 |
H46(0.00); H520(0.00) | LDD2235 | [5] | |
|
m-APA Probe Info |
 |
H46(0.00); H520(0.00) | LDD2231 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [6] | |
|
NAIA_4 Probe Info |
 |
C96(0.00); C138(0.00); C356(0.00) | LDD2226 | [7] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [8] | |
|
Acrolein Probe Info |
 |
C356(0.00); C96(0.00); H511(0.00) | LDD0217 | [9] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [7] | |
|
HHS-465 Probe Info |
 |
Y67(0.00); K71(0.00) | LDD2240 | [10] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C356(1.53) | LDD2152 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | C356(0.00); H372(0.00); H388(0.00); H511(0.00) | LDD0222 | [9] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C356(10.76) | LDD1702 | [4] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [9] |
| LDCM0023 | KB03 | MDA-MB-231 | C356(1.70) | LDD1701 | [4] |
| LDCM0109 | NEM | HeLa | H366(0.00); H520(0.00); H372(0.00); H511(0.00) | LDD0223 | [9] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C356(0.73) | LDD2107 | [4] |
| LDCM0518 | Nucleophilic fragment 22a | MDA-MB-231 | C356(1.08) | LDD2111 | [4] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C356(0.99) | LDD2129 | [4] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C356(0.82) | LDD2150 | [4] |
References
