Details of the Target
General Information of Target
| Target ID | LDTP11686 | |||||
|---|---|---|---|---|---|---|
| Target Name | Ras-related protein Rab-33B (RAB33B) | |||||
| Gene Name | RAB33B | |||||
| Gene ID | 83452 | |||||
| Synonyms |
Ras-related protein Rab-33B |
|||||
| 3D Structure | ||||||
| Sequence |
MSVDPMTYEAQFFGFTPQTCMLRIYIAFQDYLFEVMQAVEQVILKKLDGIPDCDISPVQI
RKCTEKFLCFMKGHFDNLFSKMEQLFLQLILRIPSNILLPEDKCKETPYSEEDFQHLQKE IEQLQEKYKTELCTKQALLAELEEQKIVQAKLKQTLTFFDELHNVGRDHGTSDFRESLVS LVQNSRKLQNIRDNVEKESKRLKIS |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Small GTPase superfamily, Rab family
|
|||||
| Subcellular location |
Golgi apparatus membrane
|
|||||
| Function | Protein transport. Acts, in coordination with RAB6A, to regulate intra-Golgi retrograde trafficking. It is involved in autophagy, acting as a modulator of autophagosome formation. | |||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C48(3.64) | LDD3331 | [1] | |
|
BTD Probe Info |
 |
C150(1.62) | LDD1700 | [2] | |
|
IA-alkyne Probe Info |
 |
C150(0.81) | LDD2185 | [3] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [4] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [4] | |
|
Phosphinate-6 Probe Info |
 |
N.A. | LDD0018 | [5] | |
|
AOyne Probe Info |
 |
12.50 | LDD0443 | [6] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [7] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
STS-2 Probe Info |
 |
N.A. | LDD0138 | [8] | |
|
DFG-out-4 Probe Info |
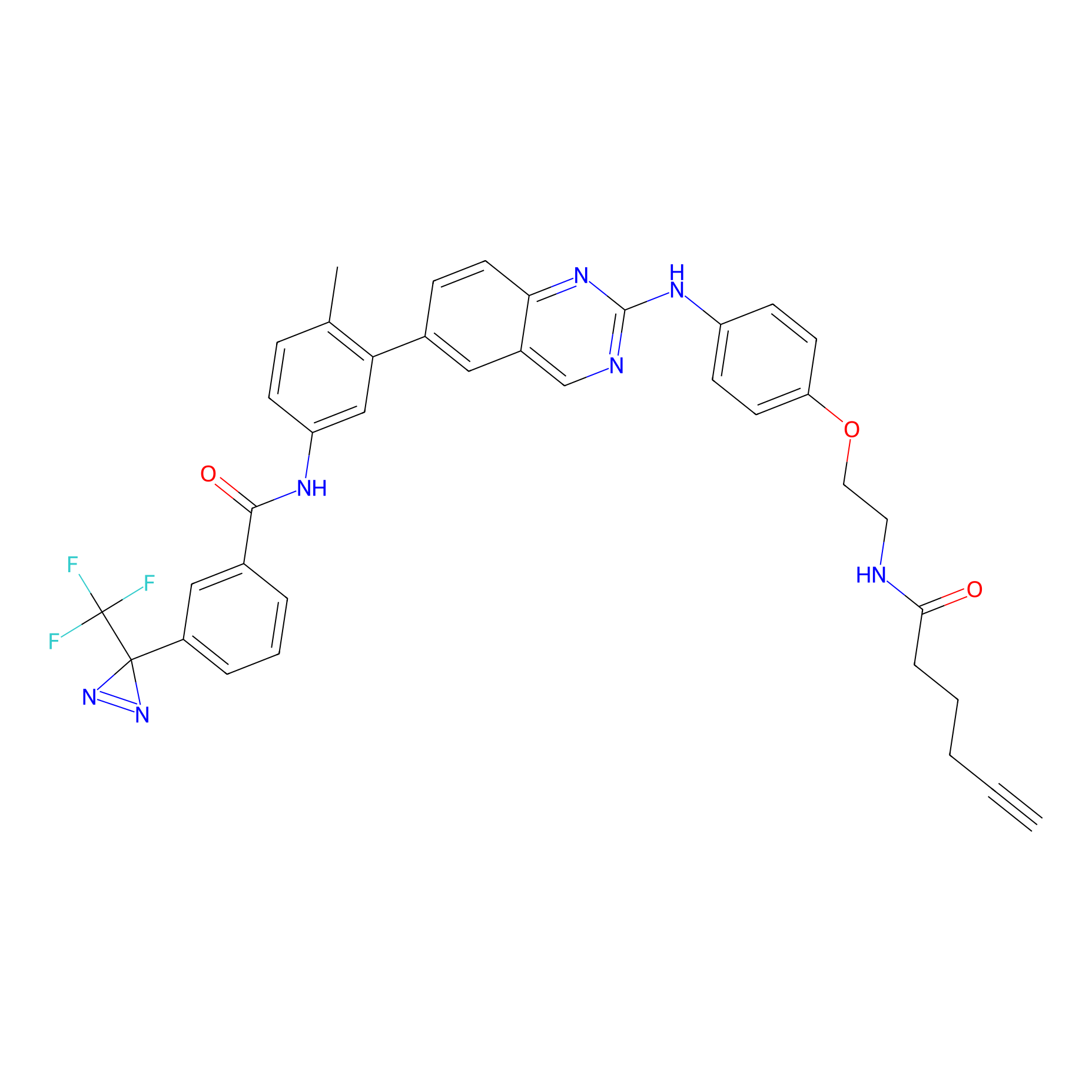 |
4.30 | LDD0075 | [9] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0524 | 2-Cyano-N-(2-morpholin-4-yl-ethyl)-acetamide | MDA-MB-231 | C150(0.71) | LDD2117 | [2] |
| LDCM0558 | 2-Cyano-N-phenylacetamide | MDA-MB-231 | C150(1.20) | LDD2152 | [2] |
| LDCM0259 | AC14 | HEK-293T | C132(1.10) | LDD1512 | [10] |
| LDCM0282 | AC22 | HEK-293T | C132(1.04) | LDD1521 | [10] |
| LDCM0291 | AC30 | HEK-293T | C132(1.09) | LDD1530 | [10] |
| LDCM0299 | AC38 | HEK-293T | C132(1.04) | LDD1538 | [10] |
| LDCM0308 | AC46 | HEK-293T | C132(0.98) | LDD1547 | [10] |
| LDCM0317 | AC54 | HEK-293T | C132(0.94) | LDD1556 | [10] |
| LDCM0323 | AC6 | HEK-293T | C132(0.86) | LDD1562 | [10] |
| LDCM0326 | AC62 | HEK-293T | C132(0.96) | LDD1565 | [10] |
| LDCM0632 | CL-Sc | Hep-G2 | C150(0.39) | LDD2227 | [7] |
| LDCM0368 | CL10 | HEK-293T | C132(1.04) | LDD1572 | [10] |
| LDCM0410 | CL22 | HEK-293T | C132(0.99) | LDD1614 | [10] |
| LDCM0423 | CL34 | HEK-293T | C132(1.04) | LDD1627 | [10] |
| LDCM0436 | CL46 | HEK-293T | C132(0.96) | LDD1640 | [10] |
| LDCM0449 | CL58 | HEK-293T | C132(0.96) | LDD1652 | [10] |
| LDCM0463 | CL70 | HEK-293T | C132(1.06) | LDD1666 | [10] |
| LDCM0476 | CL82 | HEK-293T | C132(0.91) | LDD1679 | [10] |
| LDCM0489 | CL94 | HEK-293T | C132(0.89) | LDD1692 | [10] |
| LDCM0017 | DFG-out-2 | A431 | 4.30 | LDD0075 | [9] |
| LDCM0625 | F8 | Ramos | C150(0.63) | LDD2187 | [3] |
| LDCM0573 | Fragment11 | Ramos | C150(1.20) | LDD2190 | [3] |
| LDCM0586 | Fragment28 | Ramos | C150(0.91) | LDD2198 | [3] |
| LDCM0569 | Fragment7 | Ramos | C150(0.61) | LDD2186 | [3] |
| LDCM0022 | KB02 | CAL-78 | C48(1.68) | LDD2291 | [1] |
| LDCM0023 | KB03 | MDA-MB-231 | C48(0.56) | LDD1701 | [2] |
| LDCM0024 | KB05 | MKN-1 | C48(3.64) | LDD3331 | [1] |
| LDCM0500 | Nucleophilic fragment 13a | MDA-MB-231 | C150(0.49) | LDD2093 | [2] |
| LDCM0506 | Nucleophilic fragment 16a | MDA-MB-231 | C150(0.72) | LDD2099 | [2] |
| LDCM0514 | Nucleophilic fragment 20a | MDA-MB-231 | C150(0.79) | LDD2107 | [2] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C150(0.69) | LDD2109 | [2] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C150(0.87) | LDD2119 | [2] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C150(0.53) | LDD2125 | [2] |
| LDCM0536 | Nucleophilic fragment 31 | MDA-MB-231 | C150(0.80) | LDD2129 | [2] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C150(0.58) | LDD2135 | [2] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C150(0.61) | LDD2137 | [2] |
| LDCM0211 | Nucleophilic fragment 3b | MDA-MB-231 | C150(1.62) | LDD1700 | [2] |
The Interaction Atlas With This Target
References
