Details of the Target
General Information of Target
| Target ID | LDTP09931 | |||||
|---|---|---|---|---|---|---|
| Target Name | Mitogen-activated protein kinase kinase kinase kinase 1 (MAP4K1) | |||||
| Gene Name | MAP4K1 | |||||
| Gene ID | 11184 | |||||
| Synonyms |
HPK1; Mitogen-activated protein kinase kinase kinase kinase 1; EC 2.7.11.1; Hematopoietic progenitor kinase; MAPK/ERK kinase kinase kinase 1; MEK kinase kinase 1; MEKKK 1 |
|||||
| 3D Structure | ||||||
| Sequence |
MADSKEGVLPLTAASTAPISFGFTRTSARRRLADSGDGAGPSPEEKDFLKTVEGRELQSV
KPQEAPKELVIPLIQNGHRRQPPARPPGPSTDTGALADGVVSQAVKELIAESKKSLEERE NAGVDPTLAIPMIQKGCTPSGEGADSEPRAETVPEEANYEAVPVEAYGLAMLRGMGWKPG EGIGRTFNQVVKPRVNSLRPKGLGLGANLTEAQALTPTGPSRMPRPDEEQEKDKEDQPQG LVPGGAVVVLSGPHRGLYGKVEGLDPDNVRAMVRLAVGSRVVTVSEYYLRPVSQQEFDKN TLDLRQQNGTASSRKTLWNQELYIQQDNSERKRKHLPDRQDGPAAKSEKAAPRSQHWLHR DLRVRFVDNMYKGGQYYNTKMIIEDVLSPDTCVCRTDEGRVLEGLREDMLETLVPKAEGD RVMVVLGPQTGRVGHLLSRDRARSRALVQLPRENQVVELHYDAICQYMGPSDTDDD |
|||||
| Target Type |
Clinical trial
|
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Protein kinase superfamily, STE Ser/Thr protein kinase family, STE20 subfamily
|
|||||
| Function |
Serine/threonine-protein kinase, which may play a role in the response to environmental stress. Appears to act upstream of the JUN N-terminal pathway. May play a role in hematopoietic lineage decisions and growth regulation. Able to autophosphorylate. Together with CLNK, it enhances CD3-triggered activation of T-cells and subsequent IL2 production.
|
|||||
| TTD ID | ||||||
| Uniprot ID | ||||||
| DrugMap ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
CAmbiguous(92.44); C484(7.50) | LDD0209 | [1] | |
|
CY-1 Probe Info |
 |
N.A. | LDD0246 | [2] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C428(0.00); C698(0.00); C484(0.00); C494(0.00) | LDD0038 | [3] | |
|
IA-alkyne Probe Info |
 |
C698(0.00); C494(0.00); C674(0.00); C428(0.00) | LDD0036 | [3] | |
|
Lodoacetamide azide Probe Info |
 |
C698(0.00); C484(0.00); C494(0.00); C674(0.00) | LDD0037 | [3] | |
|
KY-26 Probe Info |
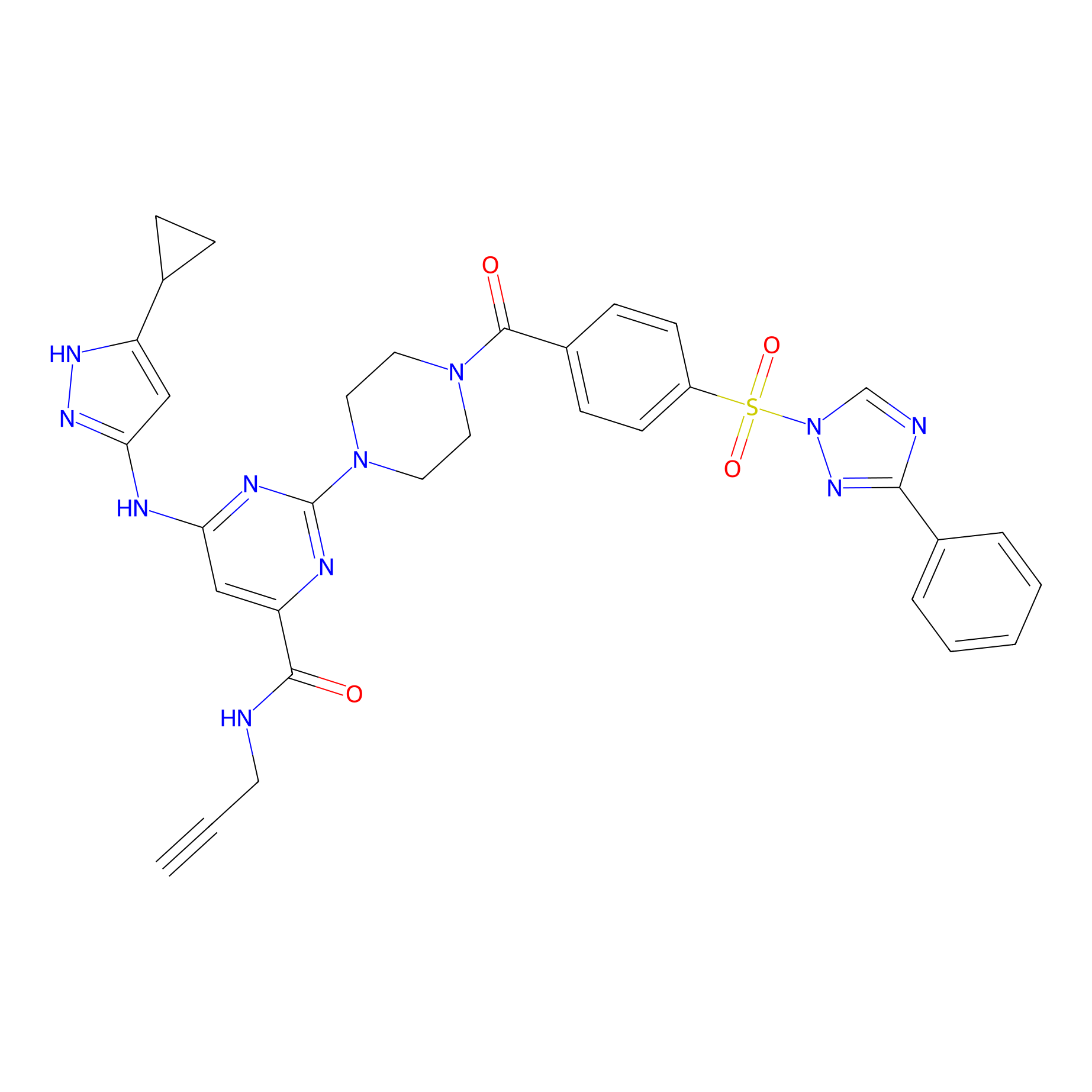 |
N.A. | LDD0301 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0625 | F8 | Ramos | C484(1.07); C428(1.74) | LDD2187 | [5] |
| LDCM0572 | Fragment10 | Ramos | C484(1.29); C428(1.24) | LDD2189 | [5] |
| LDCM0573 | Fragment11 | Ramos | C484(5.73); C428(5.96) | LDD2190 | [5] |
| LDCM0574 | Fragment12 | Ramos | C484(0.78); C428(0.87) | LDD2191 | [5] |
| LDCM0575 | Fragment13 | Ramos | C484(0.86); C428(1.10) | LDD2192 | [5] |
| LDCM0576 | Fragment14 | Ramos | C484(1.20); C428(1.15) | LDD2193 | [5] |
| LDCM0580 | Fragment21 | Ramos | C484(0.55); C428(1.03) | LDD2195 | [5] |
| LDCM0582 | Fragment23 | Ramos | C484(0.75); C428(1.07) | LDD2196 | [5] |
| LDCM0578 | Fragment27 | Ramos | C484(0.71); C428(1.20) | LDD2197 | [5] |
| LDCM0586 | Fragment28 | Ramos | C484(0.75); C428(0.72) | LDD2198 | [5] |
| LDCM0588 | Fragment30 | Ramos | C484(1.03); C428(1.37) | LDD2199 | [5] |
| LDCM0589 | Fragment31 | Ramos | C484(1.02); C428(1.22) | LDD2200 | [5] |
| LDCM0590 | Fragment32 | Ramos | C484(1.52); C428(2.71) | LDD2201 | [5] |
| LDCM0468 | Fragment33 | Ramos | C484(0.91); C428(1.12) | LDD2202 | [5] |
| LDCM0596 | Fragment38 | Ramos | C484(0.70); C428(2.62) | LDD2203 | [5] |
| LDCM0566 | Fragment4 | Ramos | C484(1.11); C428(1.13) | LDD2184 | [5] |
| LDCM0610 | Fragment52 | Ramos | C484(0.64); C428(1.72) | LDD2204 | [5] |
| LDCM0614 | Fragment56 | Ramos | C484(0.89); C428(1.91) | LDD2205 | [5] |
| LDCM0569 | Fragment7 | Ramos | C484(1.02); C428(0.93) | LDD2186 | [5] |
| LDCM0571 | Fragment9 | Ramos | C484(1.19); C428(1.63) | LDD2188 | [5] |
| LDCM0022 | KB02 | T cell | C484(9.35) | LDD1703 | [6] |
| LDCM0023 | KB03 | Jurkat | CAmbiguous(92.44); C484(7.50) | LDD0209 | [1] |
| LDCM0024 | KB05 | MONO-MAC-6 | C494(2.41) | LDD3335 | [7] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Heat shock protein HSP 90-beta (HSP90AB1) | Heat shock protein 90 family | P08238 | |||
Immunoglobulin
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Signaling lymphocytic activation molecule (SLAMF1) | . | Q13291 | |||
Other
References
