Details of the Target
General Information of Target
| Target ID | LDTP08475 | |||||
|---|---|---|---|---|---|---|
| Target Name | Carabin (TBC1D10C) | |||||
| Gene Name | TBC1D10C | |||||
| Gene ID | 374403 | |||||
| Synonyms |
Carabin; TBC1 domain family member 10C |
|||||
| 3D Structure | ||||||
| Sequence |
MAQALGEDLVQPPELQDDSSSLGSDSELSGPGPYRQADRYGFIGGSSAEPGPGHPPADLI
RQREMKWVEMTSHWEKTMSRRYKKVKMQCRKGIPSALRARCWPLLCGAHVCQKNSPGTYQ ELAEAPGDPQWMETIGRDLHRQFPLHEMFVSPQGHGQQGLLQVLKAYTLYRPEQGYCQAQ GPVAAVLLMHLPPEEAFWCLVQICEVYLPGYYGPHMEAVRLDAEVFMALLRRLLPHVHKH LQQVGVGPLLYLPEWFLCLFARSLPFPTVLRVWDAFLSEGARVLFRVGLTLVRLALGTAE QRGACPGLLETLGALRAIPPAQLQEEAFMSQVHSVVLSERDLQREIKAQLAQLPDSAPGP PPRPQVRLAGAQAIFEAQQLAGVRRGAKPEVPRIVVQPPEEPRPPRRKPQTRGKTFHGLL TRARGPPIEGPPRPQRGSTSFLDTRF |
|||||
| Target Bioclass |
Other
|
|||||
| Function |
Inhibits the Ras signaling pathway through its intrinsic Ras GTPase-activating protein (GAP) activity. Acts as a negative feedback inhibitor of the calcineurin signaling pathway that also mediates crosstalk between calcineurin and Ras.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| 22RV1 | SNV: p.R384Ter | . | |||
| HCT116 | Deletion: p.I428SfsTer126 | . | |||
| HCT15 | Substitution: p.V201A | . | |||
| HT1197 | SNV: p.M70I | . | |||
| IGR1 | SNV: p.A218V | . | |||
| JVM3 | SNV: p.Y119H | . | |||
| MOLT4 | Deletion: p.R433GfsTer121 SNV: p.E205K;p.G386C |
IA-alkyne Probe Info | |||
| NCIH1155 | SNV: p.P33T | . | |||
| NCIH2170 | SNV: p.H238L | . | |||
| OV56 | SNV: p.S29I | . | |||
| SAOS2 | SNV: p.P55L | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DBIA Probe Info |
 |
C305(1.14) | LDD3338 | [1] | |
|
YY4-yne Probe Info |
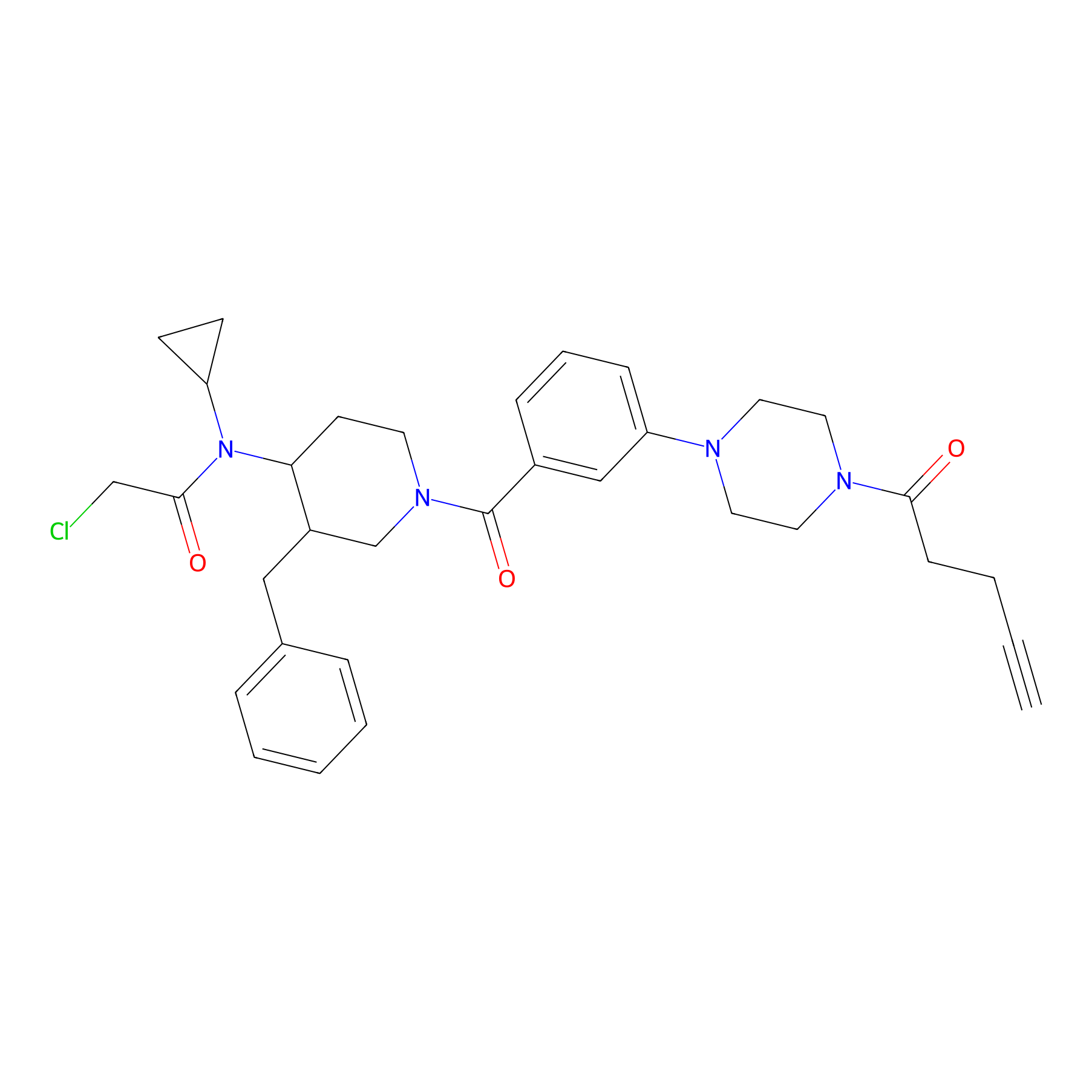 |
2.75 | LDD0400 | [2] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0166 | [3] | |
|
Lodoacetamide azide Probe Info |
 |
N.A. | LDD0037 | [4] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0625 | F8 | Ramos | C305(0.79) | LDD2187 | [5] |
| LDCM0572 | Fragment10 | Ramos | C305(0.72) | LDD2189 | [5] |
| LDCM0573 | Fragment11 | Ramos | C305(8.30) | LDD2190 | [5] |
| LDCM0574 | Fragment12 | Ramos | C305(1.29) | LDD2191 | [5] |
| LDCM0575 | Fragment13 | Ramos | C305(1.05) | LDD2192 | [5] |
| LDCM0576 | Fragment14 | Ramos | C305(1.05) | LDD2193 | [5] |
| LDCM0579 | Fragment20 | Ramos | C305(0.89) | LDD2194 | [5] |
| LDCM0580 | Fragment21 | Ramos | C305(0.84) | LDD2195 | [5] |
| LDCM0582 | Fragment23 | Ramos | C305(0.47) | LDD2196 | [5] |
| LDCM0578 | Fragment27 | Ramos | C305(1.04) | LDD2197 | [5] |
| LDCM0586 | Fragment28 | Ramos | C305(0.80) | LDD2198 | [5] |
| LDCM0588 | Fragment30 | Ramos | C305(1.56) | LDD2199 | [5] |
| LDCM0590 | Fragment32 | Ramos | C305(0.81) | LDD2201 | [5] |
| LDCM0468 | Fragment33 | Ramos | C305(1.05) | LDD2202 | [5] |
| LDCM0596 | Fragment38 | Ramos | C305(0.85) | LDD2203 | [5] |
| LDCM0566 | Fragment4 | Ramos | C305(0.66) | LDD2184 | [5] |
| LDCM0610 | Fragment52 | Ramos | C305(1.16) | LDD2204 | [5] |
| LDCM0614 | Fragment56 | Ramos | C305(1.11) | LDD2205 | [5] |
| LDCM0569 | Fragment7 | Ramos | C305(0.96) | LDD2186 | [5] |
| LDCM0571 | Fragment9 | Ramos | C305(0.83) | LDD2188 | [5] |
| LDCM0022 | KB02 | Ramos | C305(0.43) | LDD2182 | [5] |
| LDCM0023 | KB03 | Ramos | C305(1.56) | LDD2183 | [5] |
| LDCM0024 | KB05 | MV4-11 | C305(1.14) | LDD3338 | [1] |
| LDCM0154 | YY4 | T cell | 2.75 | LDD0400 | [2] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Homeobox protein Hox-A1 (HOXA1) | Antp homeobox family | P49639 | |||
Other
References
