Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
m-APA Probe Info |
 |
10.80 | LDD0402 | [1] | |
|
Probe 1 Probe Info |
 |
Y117(61.11) | LDD3495 | [2] | |
|
P11 Probe Info |
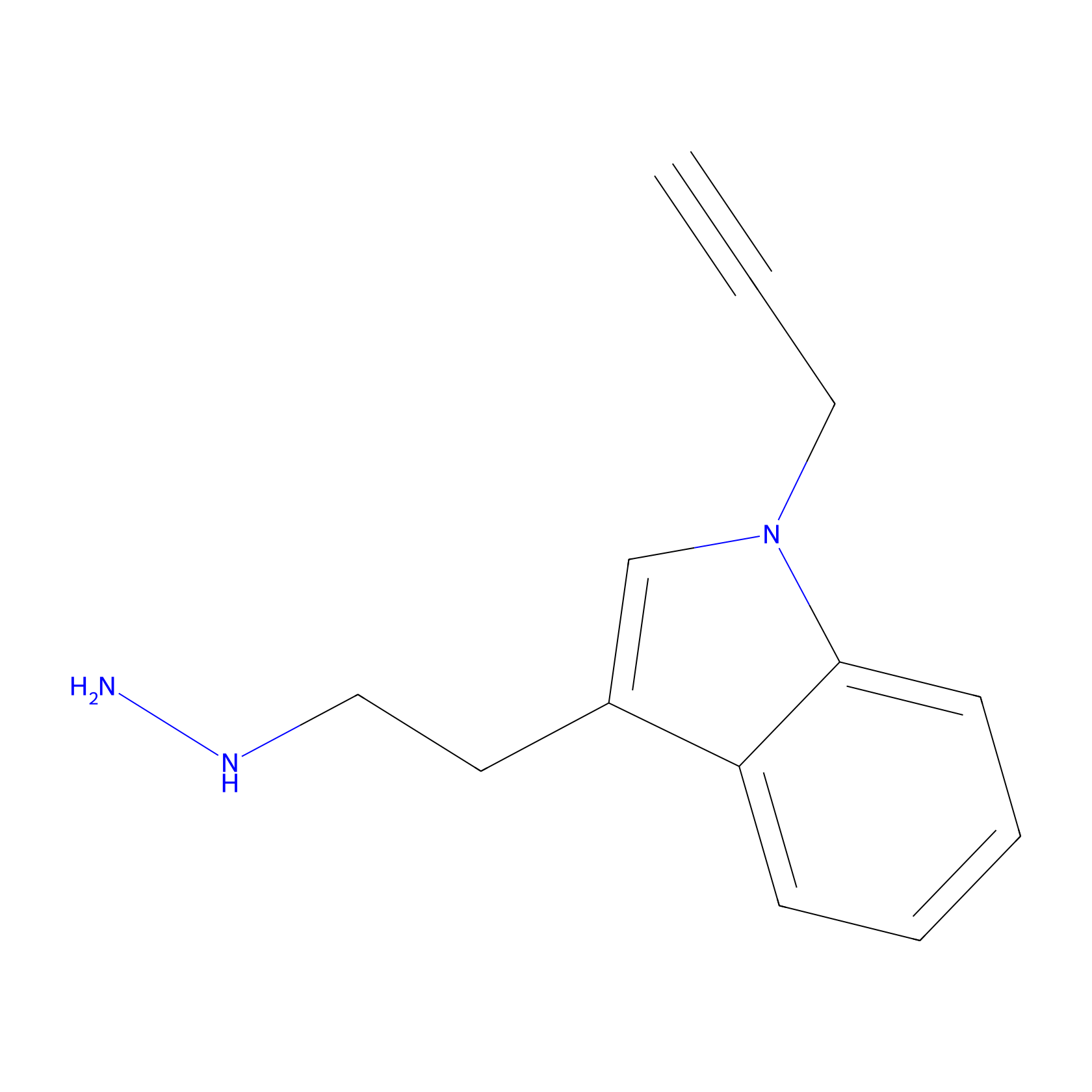 |
12.72 | LDD0201 | [3] | |
|
DBIA Probe Info |
 |
C87(0.86) | LDD3311 | [4] | |
|
HHS-465 Probe Info |
 |
Y149(10.00); Y207(10.00); Y9(10.00) | LDD2237 | [5] | |
|
ATP probe Probe Info |
 |
K86(0.00); K157(0.00); K160(0.00); K91(0.00) | LDD0199 | [6] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C71(0.00); C65(0.00) | LDD0038 | [7] | |
|
IA-alkyne Probe Info |
 |
C71(0.00); C92(0.00); C150(0.00) | LDD0032 | [8] | |
|
Lodoacetamide azide Probe Info |
 |
C71(0.00); C7(0.00); C10(0.00); C65(0.00) | LDD0037 | [7] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [9] | |
|
IPM Probe Info |
 |
C7(0.00); C71(0.00); C36(0.00); C27(0.00) | LDD0005 | [9] | |
|
SF Probe Info |
 |
N.A. | LDD0028 | [10] | |
|
STPyne Probe Info |
 |
K83(0.00); K160(0.00); K157(0.00) | LDD0009 | [9] | |
|
Phosphinate-6 Probe Info |
 |
C89(0.00); C92(0.00) | LDD0018 | [11] | |
|
Ox-W18 Probe Info |
 |
N.A. | LDD2175 | [12] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [13] | |
|
Acrolein Probe Info |
 |
C36(0.00); C65(0.00); H154(0.00) | LDD0217 | [14] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [14] | |
|
AOyne Probe Info |
 |
9.90 | LDD0443 | [15] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C7(3.02); C65(2.54) | LDD0372 | [16] |
| LDCM0156 | Aniline | NCI-H1299 | N.A. | LDD0405 | [1] |
| LDCM0020 | ARS-1620 | HCC44 | C10(1.21); C7(1.21); C209(1.05); C212(1.05) | LDD2171 | [17] |
| LDCM0088 | C45 | HEK-293T | 12.72 | LDD0201 | [3] |
| LDCM0108 | Chloroacetamide | HeLa | C65(0.00); H154(0.00); C132(0.00); C10(0.00) | LDD0222 | [14] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C71(0.71) | LDD1702 | [18] |
| LDCM0107 | IAA | HeLa | N.A. | LDD0221 | [14] |
| LDCM0022 | KB02 | 42-MG-BA | C87(1.32); C94(1.55) | LDD2244 | [4] |
| LDCM0023 | KB03 | Jurkat | C150(4.49) | LDD0315 | [8] |
| LDCM0024 | KB05 | G361 | C87(0.86) | LDD3311 | [4] |
| LDCM0109 | NEM | HeLa | H62(0.00); H73(0.00) | LDD0223 | [14] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C150(1.24); C71(1.13) | LDD2206 | [19] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C71(1.07); C150(1.04); C71(0.60) | LDD2207 | [19] |
| LDCM0131 | RA190 | MM1.R | C132(1.35); C71(1.15); C188(1.03); C191(1.03) | LDD0304 | [20] |
| LDCM0021 | THZ1 | HCT 116 | C10(1.21); C7(1.21); C209(1.05); C212(1.05) | LDD2173 | [17] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Serine/threonine-protein phosphatase 2A catalytic subunit beta isoform (PPP2CB) | PPP phosphatase family | P62714 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Major prion protein (PRNP) | Prion family | P04156 | |||
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Nuclear receptor-interacting protein 1 (NRIP1) | . | P48552 | |||
References
