Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
DA-P2 Probe Info |
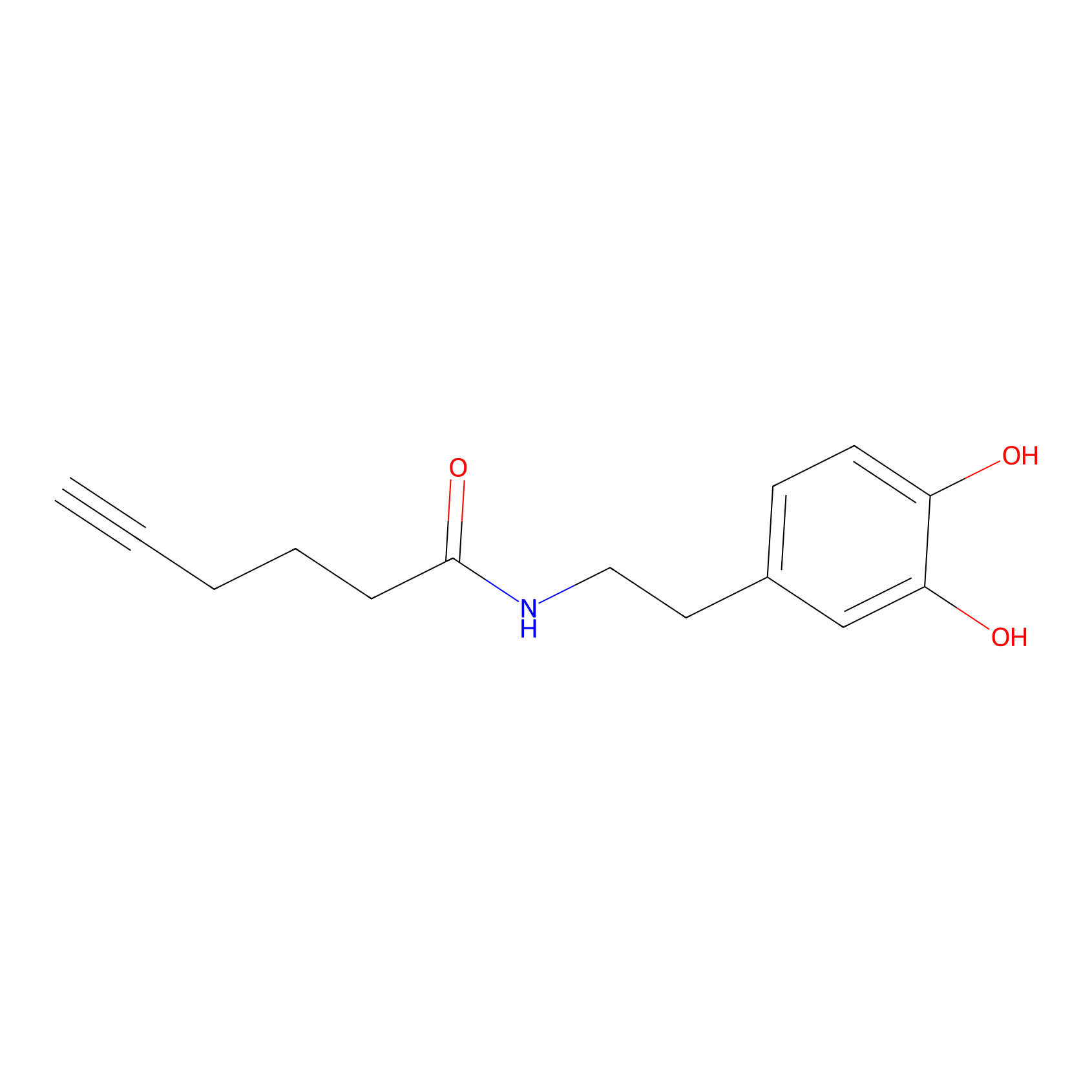 |
6.44 | LDD0348 | [1] | |
|
STPyne Probe Info |
 |
K57(0.70) | LDD0277 | [2] | |
|
m-APA Probe Info |
 |
14.08 | LDD0403 | [3] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [4] | |
|
ATP probe Probe Info |
 |
N.A. | LDD0199 | [5] | |
|
TFBX Probe Info |
 |
N.A. | LDD0027 | [6] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [7] | |
|
ENE Probe Info |
 |
C7(0.00); C62(0.00) | LDD0006 | [7] | |
|
IA-alkyne Probe Info |
 |
N.A. | LDD0151 | [8] | |
|
IPM Probe Info |
 |
C62(0.00); C7(0.00) | LDD0005 | [7] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [7] | |
|
Phosphinate-6 Probe Info |
 |
C62(0.00); C7(0.00) | LDD0018 | [9] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [10] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [11] | |
|
Crotonaldehyde Probe Info |
 |
N.A. | LDD0219 | [11] | |
|
Methacrolein Probe Info |
 |
N.A. | LDD0218 | [11] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2223 | [12] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0156 | Aniline | NCI-H1299 | 14.08 | LDD0403 | [3] |
| LDCM0108 | Chloroacetamide | HeLa | C62(0.00); H97(0.00) | LDD0222 | [11] |
| LDCM0632 | CL-Sc | Hep-G2 | C7(1.14) | LDD2227 | [12] |
| LDCM0634 | CY-0357 | Hep-G2 | C7(0.65) | LDD2228 | [12] |
| LDCM0107 | IAA | HeLa | H97(0.00); C62(0.00) | LDD0221 | [11] |
| LDCM0109 | NEM | HeLa | N.A. | LDD0223 | [11] |
| LDCM0627 | NUDT7-COV-1 | HEK-293T | C7(0.98) | LDD2206 | [13] |
| LDCM0628 | OTUB2-COV-1 | HEK-293T | C7(0.55) | LDD2207 | [13] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Protein argonaute-2 (AGO2) | Argonaute family | Q9UKV8 | |||
| Serine/threonine-protein kinase mTOR (MTOR) | PI3/PI4-kinase family | P42345 | |||
Transcription factor
Other
References
