Details of the Target
General Information of Target
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K22(2.30); K31(0.31); K35(0.44); K48(1.01) | LDD0274 | [1] | |
|
STPyne Probe Info |
 |
K18(9.70); K22(5.33); K26(6.29); K28(3.41) | LDD0277 | [1] | |
|
OPA-S-S-alkyne Probe Info |
 |
K57(3.85); K48(5.05); K35(6.43) | LDD3494 | [2] | |
|
5E-2FA Probe Info |
 |
N.A. | LDD2235 | [3] | |
|
m-APA Probe Info |
 |
N.A. | LDD2231 | [3] | |
|
CY-1 Probe Info |
 |
N.A. | LDD0246 | [4] | |
|
AZ-9 Probe Info |
 |
N.A. | LDD0395 | [5] | |
|
IPM Probe Info |
 |
N.A. | LDD0025 | [6] | |
|
JW-RF-010 Probe Info |
 |
N.A. | LDD0026 | [6] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [7] | |
|
NAIA_5 Probe Info |
 |
N.A. | LDD2224 | [7] | |
|
WYneN Probe Info |
 |
N.A. | LDD0021 | [8] | |
|
IA-alkyne Probe Info |
 |
C86(0.00); C81(0.00); C76(0.00) | LDD0151 | [9] | |
|
TFBX Probe Info |
 |
N.A. | LDD0148 | [6] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0217 | [10] | |
|
HHS-465 Probe Info |
 |
K22(0.00); K35(0.00); K31(0.00); K48(0.00) | LDD2240 | [11] | |
PAL-AfBPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
KIRA6 AfBP Probe Info |
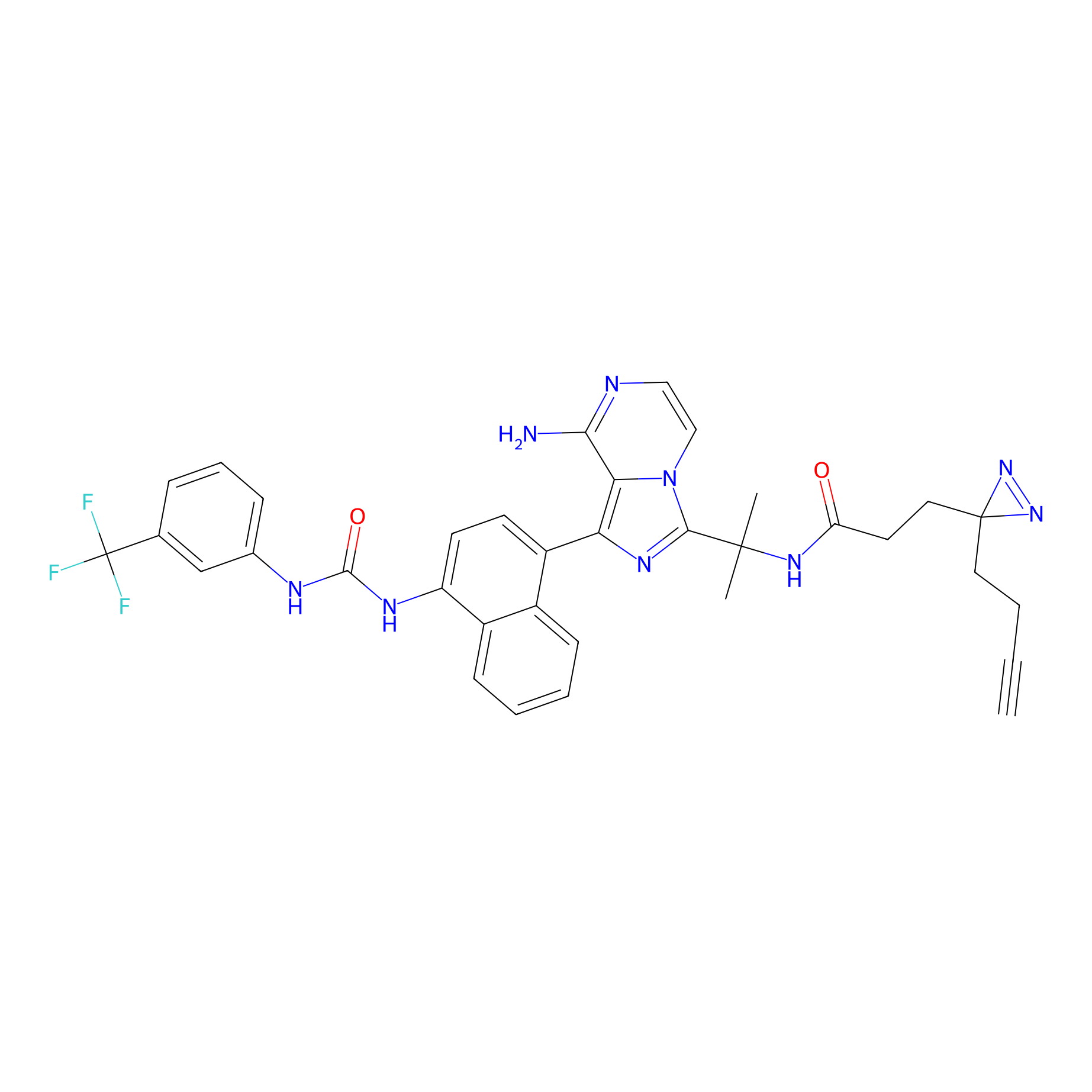 |
N.A. | LDD0378 | [12] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
Transporter and channel
Other
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Amphiregulin (AREG) | Amphiregulin family | P15514 | |||
| Protein S100-A4 (S100A4) | S-100 family | P26447 | |||
| Ezrin (EZR) | . | P15311 | |||
| Probetacellulin (BTC) | . | P35070 | |||
References
