Details of the Target
General Information of Target
| Target ID | LDTP01036 | |||||
|---|---|---|---|---|---|---|
| Target Name | Leupaxin (LPXN) | |||||
| Gene Name | LPXN | |||||
| Gene ID | 9404 | |||||
| Synonyms |
LDLP; Leupaxin |
|||||
| 3D Structure | ||||||
| Sequence |
MEELDALLEELERSTLQDSDEYSNPAPLPLDQHSRKETNLDETSEILSIQDNTSPLPAQL
VYTTNIQELNVYSEAQEPKESPPPSKTSAAAQLDELMAHLTEMQAKVAVRADAGKKHLPD KQDHKASLDSMLGGLEQELQDLGIATVPKGHCASCQKPIAGKVIHALGQSWHPEHFVCTH CKEEIGSSPFFERSGLAYCPNDYHQLFSPRCAYCAAPILDKVLTAMNQTWHPEHFFCSHC GEVFGAEGFHEKDKKPYCRKDFLAMFSPKCGGCNRPVLENYLSAMDTVWHPECFVCGDCF TSFSTGSFFELDGRPFCELHYHHRRGTLCHGCGQPITGRCISAMGYKFHPEHFVCAFCLT QLSKGIFREQNDKTYCQPCFNKLFPL |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
Paxillin family
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Transcriptional coactivator for androgen receptor (AR) and serum response factor (SRF). Contributes to the regulation of cell adhesion, spreading and cell migration and acts as a negative regulator in integrin-mediated cell adhesion events. Suppresses the integrin-induced tyrosine phosphorylation of paxillin (PXN). May play a critical role as an adapter protein in the formation of the adhesion zone in osteoclasts. Negatively regulates B-cell antigen receptor (BCR) signaling.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Target Site Mutations in Different Cell Lines
| Cell line | Mutation details | Probe for labeling this protein in this cell | |||
|---|---|---|---|---|---|
| NCIH1703 | SNV: p.H151N | DBIA Probe Info | |||
| NCIH2286 | SNV: p.E68Ter | . | |||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y375(20.00) | LDD0260 | [1] | |
|
DBIA Probe Info |
 |
C204(0.92) | LDD3310 | [2] | |
|
MCL-4 Probe Info |
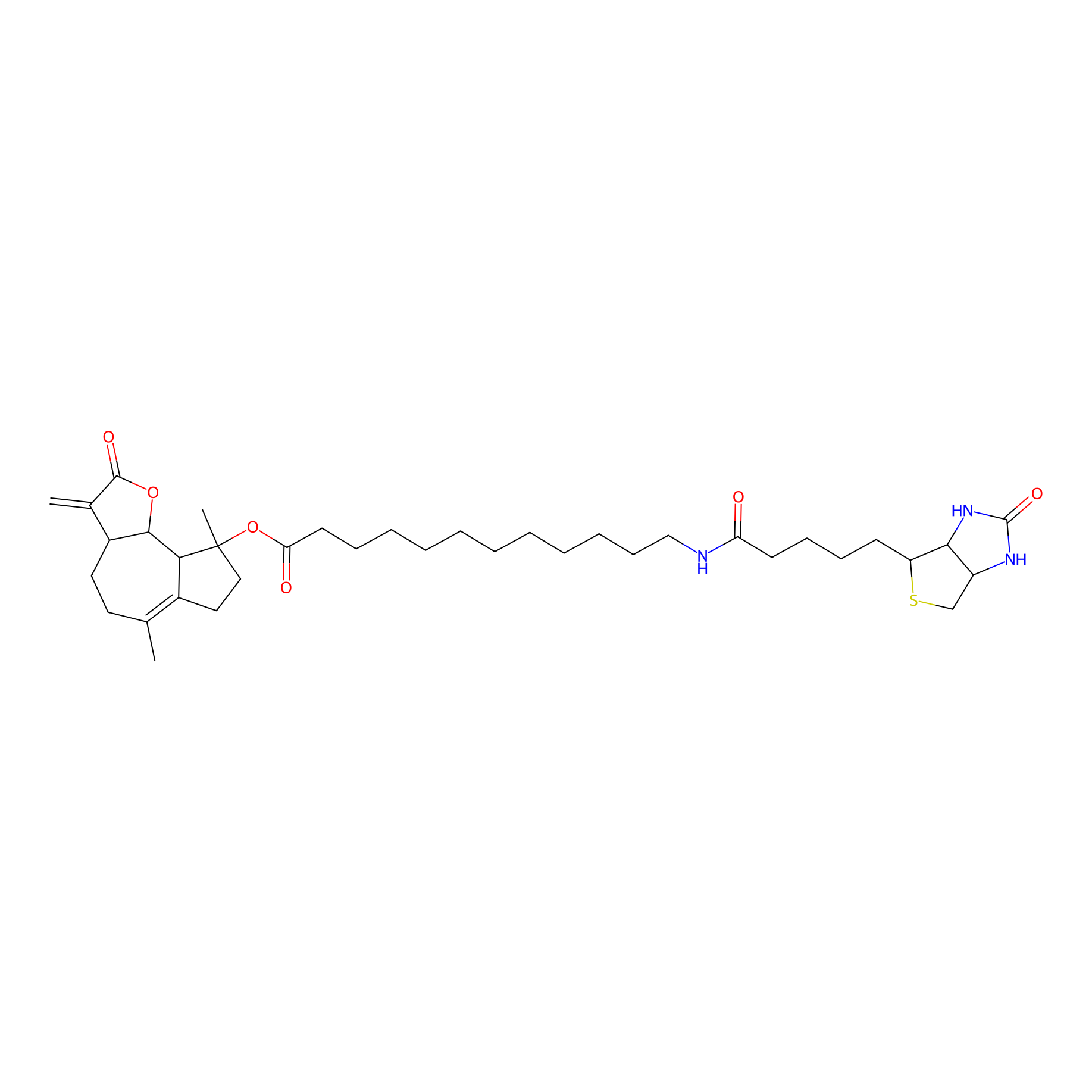 |
2.50 | LDD0049 | [3] | |
|
HHS-475 Probe Info |
 |
Y203(1.10) | LDD0264 | [4] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C340(0.00); C199(0.00); C358(0.00); C355(0.00) | LDD0038 | [5] | |
|
IA-alkyne Probe Info |
 |
C199(0.00); C340(0.00); C358(0.00); C355(0.00) | LDD0036 | [5] | |
|
Lodoacetamide azide Probe Info |
 |
C199(0.00); C340(0.00); C258(0.00); C358(0.00) | LDD0037 | [5] | |
|
NAIA_4 Probe Info |
 |
N.A. | LDD2226 | [6] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0625 | F8 | Ramos | C199(1.11); C258(2.95); C340(1.22) | LDD2187 | [7] |
| LDCM0572 | Fragment10 | Ramos | C199(0.55); C340(0.78) | LDD2189 | [7] |
| LDCM0573 | Fragment11 | Ramos | 0.47; C199(0.02); C340(0.10) | LDD2190 | [7] |
| LDCM0574 | Fragment12 | Ramos | C199(0.63); C340(0.88) | LDD2191 | [7] |
| LDCM0575 | Fragment13 | Ramos | C199(1.07); C340(1.44) | LDD2192 | [7] |
| LDCM0576 | Fragment14 | Ramos | 0.69; C199(0.80); C258(0.76); C340(0.99) | LDD2193 | [7] |
| LDCM0579 | Fragment20 | Ramos | C199(0.55); C340(0.87) | LDD2194 | [7] |
| LDCM0580 | Fragment21 | Ramos | C199(0.96); C340(1.03) | LDD2195 | [7] |
| LDCM0582 | Fragment23 | Ramos | C199(1.09); C340(1.20) | LDD2196 | [7] |
| LDCM0578 | Fragment27 | Ramos | C199(1.06); C340(1.15) | LDD2197 | [7] |
| LDCM0586 | Fragment28 | Ramos | 0.94; C199(0.81); C258(0.73); C340(5.30) | LDD2198 | [7] |
| LDCM0588 | Fragment30 | Ramos | C199(0.94); C340(1.51) | LDD2199 | [7] |
| LDCM0589 | Fragment31 | Ramos | C199(1.06); C340(0.80) | LDD2200 | [7] |
| LDCM0590 | Fragment32 | Ramos | C199(0.52); C340(0.75) | LDD2201 | [7] |
| LDCM0468 | Fragment33 | Ramos | C199(0.84); C340(0.99) | LDD2202 | [7] |
| LDCM0596 | Fragment38 | Ramos | C199(0.81); C340(1.03) | LDD2203 | [7] |
| LDCM0566 | Fragment4 | Ramos | C199(0.67); C258(0.74); C340(0.83) | LDD2184 | [7] |
| LDCM0610 | Fragment52 | Ramos | C199(1.13) | LDD2204 | [7] |
| LDCM0614 | Fragment56 | Ramos | C199(1.11); C340(1.30) | LDD2205 | [7] |
| LDCM0569 | Fragment7 | Ramos | 0.32; C199(0.49); C258(0.67); C340(0.67) | LDD2186 | [7] |
| LDCM0571 | Fragment9 | Ramos | C199(0.53); C340(0.93) | LDD2188 | [7] |
| LDCM0116 | HHS-0101 | DM93 | Y203(1.10) | LDD0264 | [4] |
| LDCM0117 | HHS-0201 | DM93 | Y203(0.19) | LDD0265 | [4] |
| LDCM0118 | HHS-0301 | DM93 | Y203(0.67) | LDD0266 | [4] |
| LDCM0120 | HHS-0701 | DM93 | Y203(0.65) | LDD0268 | [4] |
| LDCM0022 | KB02 | Ramos | 0.23; C199(0.44); C258(0.52); C340(0.75) | LDD2182 | [7] |
| LDCM0023 | KB03 | Jurkat | C340(10.90) | LDD0315 | [8] |
| LDCM0024 | KB05 | COLO792 | C204(0.92) | LDD3310 | [2] |
| LDCM0006 | Micheliolide | M9-ENL1 | 2.50 | LDD0049 | [3] |
| LDCM0131 | RA190 | MM1.R | C340(1.28) | LDD0304 | [9] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Alanine--glyoxylate aminotransferase (AGXT) | Class-V pyridoxal-phosphate-dependent aminotransferase family | P21549 | |||
| Tensin-2 (TNS2) | PTEN phosphatase protein family | Q63HR2 | |||
Transporter and channel
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Aquaporin-1 (AQP1) | MIP/aquaporin (TC 1.A.8) family | P29972 | |||
| Optineurin (OPTN) | . | Q96CV9 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Homeobox protein Hox-C9 (HOXC9) | Abd-B homeobox family | P31274 | |||
Cytokine and receptor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transforming growth factor beta-1 proprotein (TGFB1) | TGF-beta family | P01137 | |||
Other
References
