Details of the Target
General Information of Target
| Target ID | LDTP09213 | |||||
|---|---|---|---|---|---|---|
| Target Name | Diphosphoinositol polyphosphate phosphohydrolase 3-alpha (NUDT10) | |||||
| Gene Name | NUDT10 | |||||
| Gene ID | 170685 | |||||
| Synonyms |
APS2; DIPP3A; Diphosphoinositol polyphosphate phosphohydrolase 3-alpha; DIPP-3-alpha; DIPP3-alpha; hDIPP3alpha; EC 3.6.1.52; Diadenosine 5',5'''-P1,P6-hexaphosphate hydrolase 3-alpha; Diadenosine hexaphosphate hydrolase; AMP-forming); EC 3.6.1.60; Nucleoside diphosphate-linked moiety X motif 10; Nudix motif 10; hAps2
|
|||||
| 3D Structure | ||||||
| Sequence |
MKCKPNQTRTYDPEGFKKRAACLCFRSEREDEVLLVSSSRYPDRWIVPGGGMEPEEEPGG
AAVREVYEEAGVKGKLGRLLGVFEQNQDPKHRTYVYVLTVTELLEDWEDSVSIGRKREWF KVEDAIKVLQCHKPVHAEYLEKLKLGGSPTNGNSMAPSSPDSDP |
|||||
| Target Bioclass |
Enzyme
|
|||||
| Family |
Nudix hydrolase family, DIPP subfamily
|
|||||
| Subcellular location |
Cytoplasm
|
|||||
| Function |
Cleaves a beta-phosphate from the diphosphate groups in PP-InsP5 (diphosphoinositol pentakisphosphate), suggesting that it may play a role in signal transduction. Also able to catalyze the hydrolysis of dinucleoside oligophosphates, with Ap6A and Ap5A being the preferred substrates. The major reaction products are ADP and p4a from Ap6A and ADP and ATP from Ap5A. Also able to hydrolyze 5-phosphoribose 1-diphosphate.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
| ChEMBL ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
TH211 Probe Info |
 |
Y67(15.61); Y11(11.97) | LDD0257 | [1] | |
|
STPyne Probe Info |
 |
K133(20.00) | LDD2217 | [2] | |
|
Probe 1 Probe Info |
 |
Y67(48.83) | LDD3495 | [3] | |
|
m-APA Probe Info |
 |
10.72 | LDD0403 | [4] | |
|
BTD Probe Info |
 |
C131(1.10) | LDD2094 | [5] | |
|
AHL-Pu-1 Probe Info |
 |
C131(4.91) | LDD0168 | [6] | |
|
HHS-475 Probe Info |
 |
Y67(0.77) | LDD0264 | [7] | |
|
Acrolein Probe Info |
 |
N.A. | LDD0222 | [8] | |
|
4-Iodoacetamidophenylacetylene Probe Info |
 |
C3(0.00); C131(0.00) | LDD0038 | [9] | |
|
IA-alkyne Probe Info |
 |
C3(0.00); C131(0.00) | LDD0036 | [9] | |
|
Lodoacetamide azide Probe Info |
 |
C3(0.00); C131(0.00) | LDD0037 | [9] | |
|
ENE Probe Info |
 |
N.A. | LDD0006 | [10] | |
|
IPM Probe Info |
 |
N.A. | LDD0005 | [10] | |
|
NPM Probe Info |
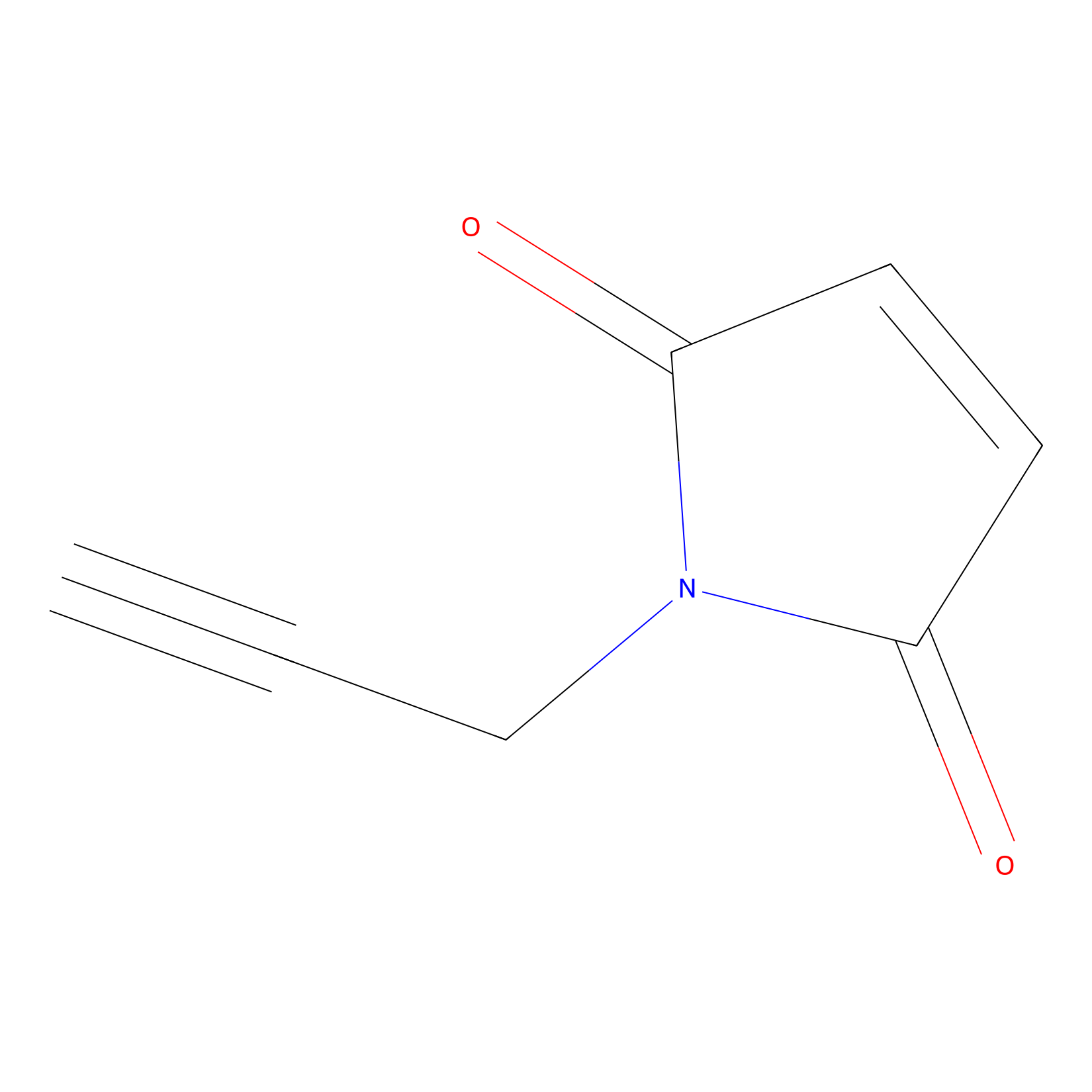 |
N.A. | LDD0016 | [10] | |
|
TFBX Probe Info |
 |
C24(0.00); C131(0.00) | LDD0148 | [11] | |
|
VSF Probe Info |
 |
N.A. | LDD0007 | [10] | |
|
Phosphinate-6 Probe Info |
 |
C3(0.00); C131(0.00) | LDD0018 | [12] | |
|
AOyne Probe Info |
 |
15.00 | LDD0443 | [13] | |
|
HHS-465 Probe Info |
 |
N.A. | LDD2240 | [14] | |
Competitor(s) Related to This Target
| Competitor ID | Name | Cell line | Binding Site(Ratio) | Interaction ID | Ref |
|---|---|---|---|---|---|
| LDCM0025 | 4SU-RNA | HEK-293T | C131(4.91) | LDD0168 | [6] |
| LDCM0026 | 4SU-RNA+native RNA | HEK-293T | C131(8.65) | LDD0169 | [6] |
| LDCM0156 | Aniline | NCI-H1299 | 10.72 | LDD0403 | [4] |
| LDCM0108 | Chloroacetamide | HeLa | N.A. | LDD0222 | [8] |
| LDCM0213 | Electrophilic fragment 2 | MDA-MB-231 | C22(1.42); C24(1.42) | LDD1702 | [5] |
| LDCM0116 | HHS-0101 | DM93 | Y67(0.77) | LDD0264 | [7] |
| LDCM0117 | HHS-0201 | DM93 | Y67(0.82) | LDD0265 | [7] |
| LDCM0118 | HHS-0301 | DM93 | Y67(0.74) | LDD0266 | [7] |
| LDCM0119 | HHS-0401 | DM93 | Y67(1.07) | LDD0267 | [7] |
| LDCM0120 | HHS-0701 | DM93 | Y67(0.90) | LDD0268 | [7] |
| LDCM0023 | KB03 | MDA-MB-231 | C24(2.21) | LDD1701 | [5] |
| LDCM0501 | Nucleophilic fragment 13b | MDA-MB-231 | C131(1.10) | LDD2094 | [5] |
| LDCM0516 | Nucleophilic fragment 21a | MDA-MB-231 | C131(1.06) | LDD2109 | [5] |
| LDCM0526 | Nucleophilic fragment 26a | MDA-MB-231 | C131(1.75) | LDD2119 | [5] |
| LDCM0527 | Nucleophilic fragment 26b | MDA-MB-231 | C22(0.87) | LDD2120 | [5] |
| LDCM0530 | Nucleophilic fragment 28a | MDA-MB-231 | C131(0.94) | LDD2123 | [5] |
| LDCM0532 | Nucleophilic fragment 29a | MDA-MB-231 | C131(0.99) | LDD2125 | [5] |
| LDCM0542 | Nucleophilic fragment 37 | MDA-MB-231 | C24(0.90) | LDD2135 | [5] |
| LDCM0544 | Nucleophilic fragment 39 | MDA-MB-231 | C131(1.10); C24(1.03) | LDD2137 | [5] |
| LDCM0549 | Nucleophilic fragment 43 | MDA-MB-231 | C24(0.94) | LDD2143 | [5] |
| LDCM0550 | Nucleophilic fragment 5a | MDA-MB-231 | C24(1.01) | LDD2144 | [5] |
| LDCM0556 | Nucleophilic fragment 8a | MDA-MB-231 | C131(0.37) | LDD2150 | [5] |
The Interaction Atlas With This Target
The Protein(s) Related To This Target
Enzyme
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| E3 ubiquitin-protein ligase TRIM8 (TRIM8) | TRIM/RBCC family | Q9BZR9 | |||
Transcription factor
| Protein name | Family | Uniprot ID | |||
|---|---|---|---|---|---|
| Transcription factor 4 (TCF4) | . | P15884 | |||
Other
References
