Details of the Target
General Information of Target
| Target ID | LDTP02102 | |||||
|---|---|---|---|---|---|---|
| Target Name | Non-histone chromosomal protein HMG-17 (HMGN2) | |||||
| Gene Name | HMGN2 | |||||
| Gene ID | 3151 | |||||
| Synonyms |
HMG17; Non-histone chromosomal protein HMG-17; High mobility group nucleosome-binding domain-containing protein 2 |
|||||
| 3D Structure | ||||||
| Sequence |
MPKRKAEGDAKGDKAKVKDEPQRRSARLSAKPAPPKPEPKPKKAPAKKGEKVPKGKKGKA
DAGKEGNNPAENGDAKTDQAQKAEGAGDAK |
|||||
| Target Bioclass |
Other
|
|||||
| Family |
HMGN family
|
|||||
| Subcellular location |
Nucleus
|
|||||
| Function |
Binds to the inner side of the nucleosomal DNA thus altering the interaction between the DNA and the histone octamer. May be involved in the process which maintains transcribable genes in a unique chromatin conformation.
|
|||||
| Uniprot ID | ||||||
| Ensemble ID | ||||||
| HGNC ID | ||||||
Probe(s) Labeling This Target
ABPP Probe
| Probe name | Structure | Binding Site(Ratio) | Interaction ID | Ref | |
|---|---|---|---|---|---|
|
ONAyne Probe Info |
 |
K31(10.00); K36(6.36); K59(0.99); K76(7.69) | LDD0274 | [1] | |
|
STPyne Probe Info |
 |
K16(2.57); K18(7.14); K31(10.00); K36(9.02) | LDD0277 | [1] | |
|
AZ-9 Probe Info |
 |
D88(4.18); D74(10.00) | LDD2209 | [2] | |
|
OPA-S-S-alkyne Probe Info |
 |
K16(1.78); K18(2.09) | LDD3494 | [3] | |
|
P11 Probe Info |
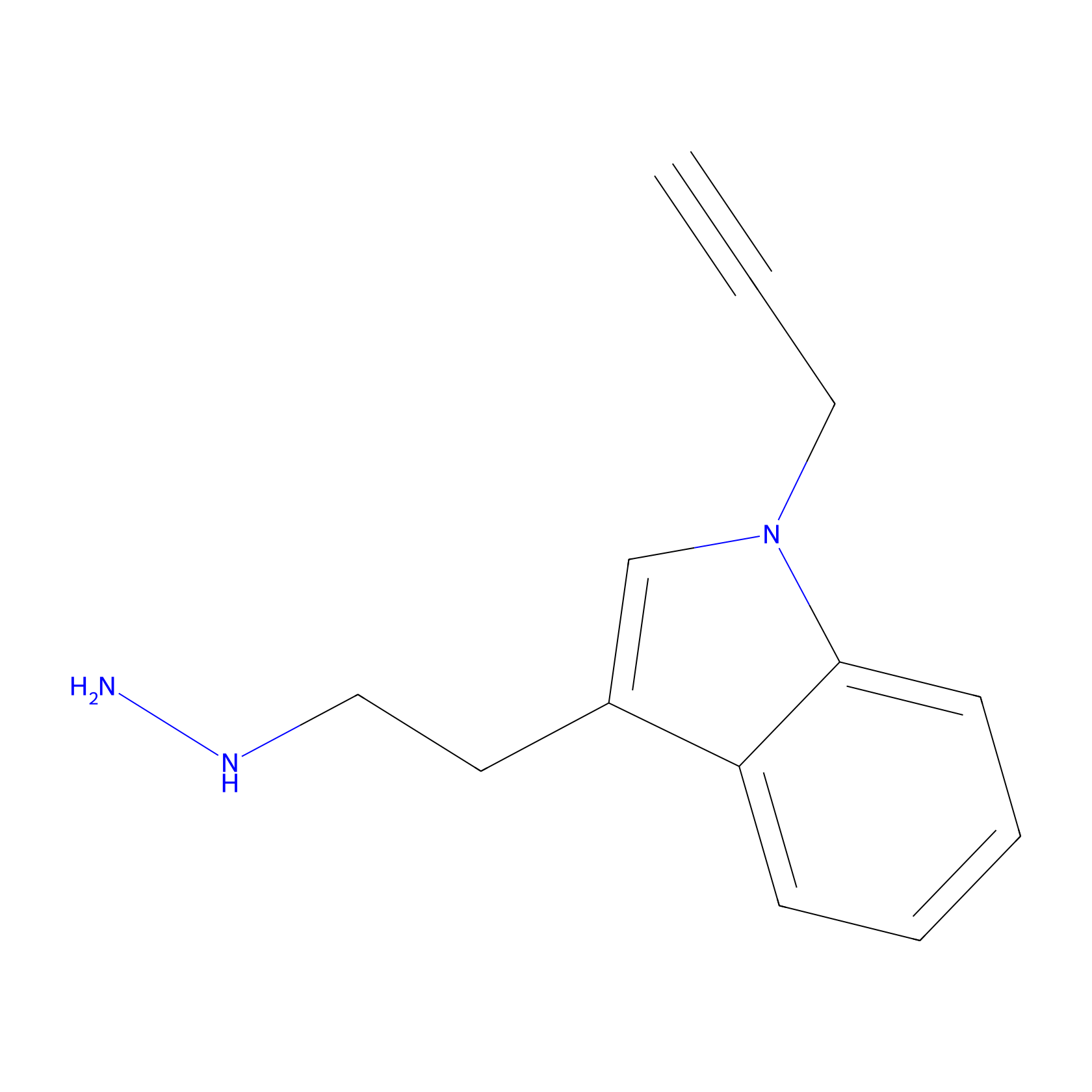 |
5.24 | LDD0201 | [4] | |
|
ATP probe Probe Info |
 |
K31(0.00); K40(0.00) | LDD0035 | [5] | |
|
NHS Probe Info |
 |
K82(0.00); K76(0.00) | LDD0010 | [6] | |
|
SF Probe Info |
 |
K11(0.00); K59(0.00); K5(0.00); K36(0.00) | LDD0028 | [7] | |
|
1c-yne Probe Info |
 |
K64(0.00); K76(0.00) | LDD0228 | [8] | |
|
1d-yne Probe Info |
 |
N.A. | LDD0357 | [8] | |
Competitor(s) Related to This Target
The Interaction Atlas With This Target
References
